Images
Participants



Contact

IRB Barcelona scientists pave the way towards describing the conformation of proteins that do not have a defined structure
Structural and theoretical techniques are combined to develop new methodologies for the analysis of proteins
Researchers with the joint program between IRB Barcelona and the Barcelona Supercomputing Center (BSC) have devised a new strategy to study the shape of proteins. This study has been led by Modesto Orozco, head of the Molecular Modeling and Bioinformatics Group, and Xavier Salvatella, head of the Molecular Biophysics Group, both ICREA scientists at IRB Barcelona. According to Orozco, also senior professor of the University of Barcelona and director of the Life Sciences Department at BSC, “by combining computational modeling and experimental physicochemical techniques, we have revealed the structures of proteins, which, until now, were unachievable because of technical barriers”. The results are available from today in the electronic version of the prestigious journal Proceedings of the National Academy of Sciences (PNAS).
Developed at IRB Barcelona, this project represents an advance in protein structure research. The first author, the Italian PhD student Michela Candotti, says, “knowing the shape that proteins have is essential to perform any analysis. A wire can be a paperclip, a staple or a spring, depending how it is folded”. This remark is especially relevant given the multi-functional nature of many proteins.
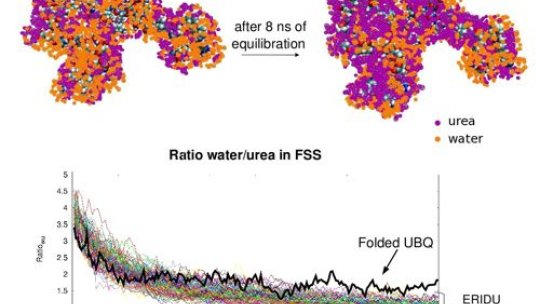
The study has several scientific implications, which can be summarized in the following three points. First of all, the researchers have described the chemical mechanisms by which compounds such as urea unfold proteins. “This was a debate that started in the 60s, and with this work it can now be considered closed”, explains Orozco. Furthermore, they have established a new strategy that will allow them to decipher the conformation of the Intrinsically Disordered Proteins (IDP). IDPs are a group of proteins without a rigid structure that comprise a large part of the proteome; however, little is known about them. “Our results will contribute to research into diseases that involve IDPs, such as cancer, Parkinson’s or Alzheimer”, asserts Salvatella. Finally, the scientists have identified the first steps in protein folding, another topic that is widely contended.
Reference article:
Towards an atomistic description of the urea-denatured state of proteins
Michela Candotti, Santiago Esteban-Martín, Xavier Salvatella and Modesto Orozco
Proceedings of the National Academy of Sciences (PNAS) (2013) online Early Edition the week of March 25.
About IRB Barcelona
The Institute for Research in Biomedicine (IRB Barcelona) pursues a society free of disease. To this end, it conducts multidisciplinary research of excellence to cure cancer and other diseases linked to ageing. It establishes technology transfer agreements with the pharmaceutical industry and major hospitals to bring research results closer to society, and organises a range of science outreach activities to engage the public in an open dialogue. IRB Barcelona is an international centre that hosts 400 researchers and more than 30 nationalities. Recognised as a Severo Ochoa Centre of Excellence since 2011, IRB Barcelona is a CERCA centre and member of the Barcelona Institute of Science and Technology (BIST).