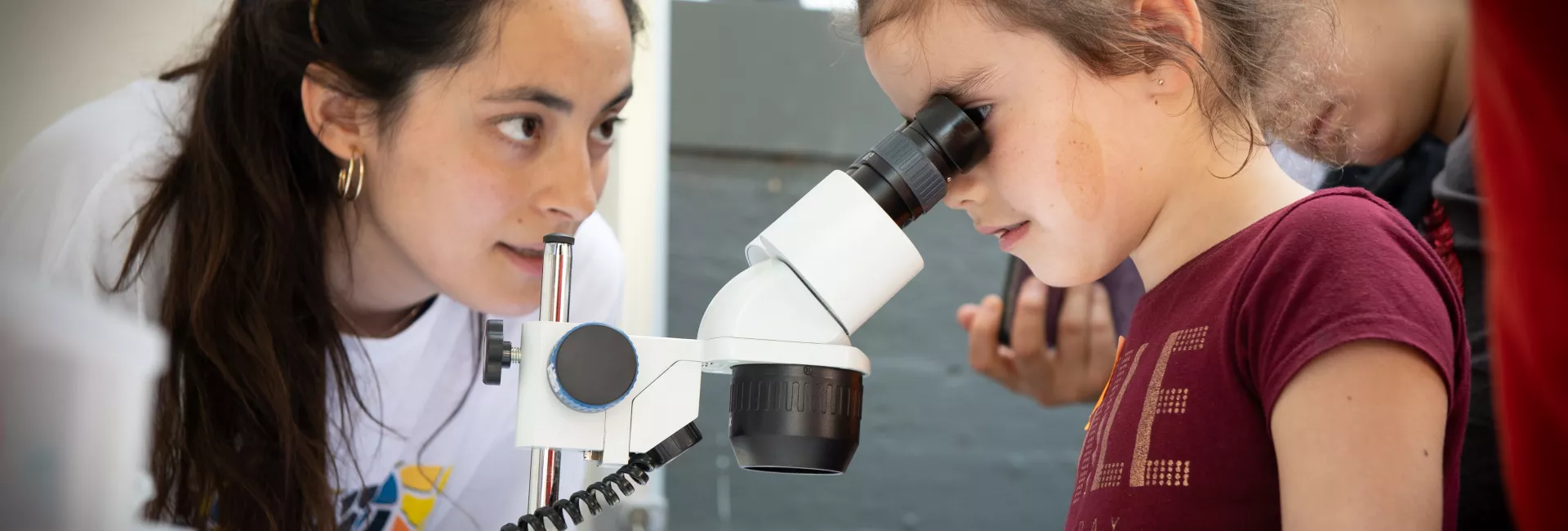
Organizado por el IRB Barcelona en colaboración con la Fundació Catalunya – La Pedrera.
Los tutores de esta edición son: Paula Climent, Anna Casas, Ricardo Viais, Najate Benhra, Begoña Cánovas, Sanja Zivanovic, Yuliana Enciso, Clara Suñer, Elzbieta Maria Szulc, Artur Ezquerra, Luis Daniel Toribio, Jorge García, Antonija Kuzmanic y Paula Martínez-Cristobal.
Presentation

Participantes del curso “Crazy About Biomedicine”, edición 2016.
Objetivos
Bojos per la Biomedicina és un curs d'un any de durada dirigit als estudiants del primer any de batxillerat que desitgin explorar alguns dels descobriments fascinants que s'estan fent actualment en les ciències de la vida. A través d'aquest curs, els estudiants tindran l'oportunitat d'aprofundir el seu coneixement de la teoria i tècniques científiques en el camp de la biomedicina. Treballaran juntament amb investigadors joves per experimentar com es fa ciència en un institut de recerca internacional, guanyar una mica d'experiència pràctica en les últimes metodologies d'avantguarda i posicionar-se per a una possible carrera professional en les ciències de la vida.
Descripción del curso
Aquest curs combina sessions teòriques i activitats experimentals pràctiques, que es duran a terme durant 18 dissabtes de l'any. El curs tractarà 12 temes científics actuals, que van des de la biologia cel·lular i molecular fins a la biologia estructural i computacional i la química, presentats per estudiants de doctorat de l'IRB Barcelona. En el primer «semestre» (gener-juny 2015), els 3 primers dissabtes es dedicaran a aquestes sessions teòriques generals per a tots els participants. Durant els 6 dissabtes següents, es formaran grups petits que entraran als laboratoris per a les sessions pràctiques. A continuació, es repetirà aquest programa amb 6 temes de recerca nous per al segon semestre (juny-desembre). Els estudiants participants s'hauran de comprometre a assistir al curs durant tot l'any.
Idioma del curso
Totes les xerrades i sessions pràctiques es faran en anglès.
Fechas y horarios
El curs es farà de gener a desembre de 2016, 10.00h-14.00h
SEMESTER I
- Divendres, 15 de gener: Inauguracióal mon Sant Benedet- Sant Fruitós del Bages
- Dissabte, 16 de gener: Seminari 1
- Dissabte, 23 de gener: Seminari 2
- Dissabte, 6 de febrer: Seminari 3
- Dissabte, 20 de febrer: Tallers 1
- Dissabte, 5 de març: Tallers 2
- Dissabte, 12 de març: Tallers 3
- Dissabte, 2 d’abril: Tallers 6
- Dissabte, 16 d’abril: Tallers 4
- Dissabte, 30 d’abril: Tallers 5
SEMESTER II
- Dissabte, 21 de maig: Seminari 1
- Dissabte, 28 de maig: Seminari 2
- Dissabte, 11 de juny: Seminari 3
- Dissabte, 17 de setembre: Tallers 1
- Dissabte, 1 d’octubre: Tallers 2
- Dissabte, 15 d’octubre: Tallers 3
- Dissabte, 29 d’octubre: Tallers 4
- Dissabte, 19 de novembre: Tallers 5
- Dissabte, 26 de novembre: Tallers 6
- Dissabte, gener de 2017: Cerimonia de cloenda
Precio del curso
Les despeses de l’acció formativa corren a càrrec de la Fundació Catalunya-La Pedrera. Tot i això els participants abonaran 120 euros corresponent a les despeses d’organització i gestió del programa. No s'inclouen dinars ni esmorzars. Els estudiants rebran un certificat de participació en finalitzar el curs amb èxit en una cerimònia especial de cloenda.
Lugar de celebración
Insitut de Recerca Biomèdica (IRB Barcelona)
c/ Baldiri Reixac, 10
Barcelona
Quién puede solicitar una plaza
Aquest curs està dirigit als estudiants de primer any de batxillerat que tinguin un interès i talent especials en els camps relacionats amb les ciències de la vida (principalment biologia i química).
Els estudiants poden sol·licitar la plaça a un màxim de 2 dels programes de la série "Bojos per la Ciència" i participar només en un.
Cómo solicitar la plaza
Els estudiants interessats hauran d'emplenar el formulari web de sol·licitud i incloure una carta de motivació. També es demanarà una carta de recomanació directament de dos dels seus professors que la coneguin bé. En el cas de que l'estudiant hagi canviat de centre aquest curs, suggerim que sol·licitin les cartes als antics professors. La data límit d'inscripció és el 23 October 2015.
El curs està obert a un total de 24 estudiants. Se seleccionaran els candidats en funció del seu expedient acadèmic, de les recomanacions dels seus professors i de la seva motivació per participar-hi. Es convidarà els candidats preseleccionats a fer entrevistes amb els organitzadors científics al novembre, després de les quals es farà la selecció final. La primera setmana de desembre es comunicarà el resultat als estudiants. Es demanarà als estudiants seleccionats per participar-hi i als seus pares/tutors legals que signin una carta de compromís d'assistir a totes les sessions.
Fechas de interés
- 23 d’octubre de 2015: Data límit d'inscripció
- Setmana del 9 de novembre de 2015: Contacte amb els candidats preseleccionats
- 16-27 de novembre de 2015: Entrevistes
- 1-4 de desembre de 2015: Publicació del llistat dels estudiants acceptats
- Divendres, 15 de gener: Inauguracióal mon Sant Benedet- Sant Fruitós del Bages
- Dissabte, 16 de gener de 2017: Inici del curs
Col·laboradors
Facebook: https://www.
Twiter: @catfundacio
Instagram: #bojosperlaciencia @catfundacio



Si tienes alguna pregunta, contactanos en: irb_outreach@irbbarcelona.org
Programme
SEMESTER 1
1. Epigenetic code: Driving fruitfly development
Paula Climent & Anna Casas (Azorin’s Lab)
Eukariote DNA is not naked but it is organized in a structure called chromatin, which is formed by nucleosomes. A nucleosome is a particle formed by core histones where DNA wraps around. During the last years, our knowledge of chromatin regulation and function has improved due to the identification of several histone modifications, histone variants, remodeling complexes and other structural and non-structural proteins. All this factors have a role in chromatin compactation and this leads to changes in gene expression. The field of epigenetics studies how these factors affect gene expression and its importance during development and diferent cell processes like replication, transcription, etc.
Our group focuses on the study of the molecular basis of chromatin structure and function and how this is regulated during the cell cycle and the development of an organism. We use Drosophila melanogaster or fruitfly as a model organism because the similarity between the fly and human genomes and for the huge amount of tools that are available.
During this course we will study one of the proteins that binds to the DNA and participates in the chromatin organization. We will focus in one of the variants of the histone H1 and some modifications to try to understand how this protein affects to the gene expression and the development of the fruitfly.
2. Study of tumorigenesis using flies
Najate Benhra (Milan’s Lab)
Cancer is a multi-hit process that involves mutations in oncogenes and tumour suppressors. It is characterised by the uncontrolled proliferation of malignant cells and invasion to adjacent healthy tissue. Aggressive cancer cells activate a programme named EMT (Epithelial to Mesenchymal Cells), which causes cancer cells to undergo morphogenetic alterations, leading to loss of their apico-basal polarity and acquisition of invasive motility. Over the last decade, the fruit fly Drosophila melanogaster has become a key model system for cancer research. This model has the advantage that the signalling pathways are conserved between flies and humans, genome redundancy is reduced compared to humans, and there are a wide range of powerful genetic tools for use in Drosophila.
In this course, we will give an overview of Drosophila genetics and present how we use this model system to characterise aspects of tumorigenesis, such as cell proliferation and cell invasion, at the molecular and cellular levels. In particular, we will focus on the “wing imaginal disc”, which will allow us to analyse the signalling pathways involved in some aspects of tumorigenesis.
3. Stepping into the world of the cell skeleton
Ricardo Viais (Lüder’s Lab)
The cytoskeleton is a cellular compartment present in cells from all domains of life. As its name suggests, it is the “skeleton” of the cell, ensuring its shape and migration. In eukaryotic cells, one of the components of the cytoskeleton are microtubules—tiny tubular structures that form a network throughout the cytoplasm. Microtubules can be considered LEGO, where the basic individual bricks are monomers of a protein called tubulin. The microtubule network is extremely dynamic, and the addition or removal of tubulin “pieces” from microtubules leads to their growth or shrinkage. In fact, microtubules are much more than just a “skeleton of the cell”. They are also involved in chromosome guidance during mitosis and meiosis and they serve as highways for the transport of organelles and key molecules inside the cell. Microtubules are one of the targets of cancer treatment and have also been shown to be involved in neurodegenerative diseases (like Parkinson’s and Alzheimer’s).
In this practical course, we will learn about how microtubules are formed, how they grow, and what they look like in different cell types. For this purpose, we will use tissue cultured cell lines, mouse hippocampal neurons, and fluorescence light microscopy techniques.
4. Study of novel cancer treatments
Begoña Cánovas (Nebreda’s lab)
Cancer is not a disease,- but a generic way of defining a broad group of diseases characterised by the uncontrolled proliferation of a group of cells.
Homeostasis calls for the proper control of cell proliferation, so all the cells in our bodies are subjected to certain “checkpoints” that detect alterations. Upon any modification in cell behaviour, these “checkpoints” are activated and lead the “defective” cells to apoptosis.
Cancer cells, however, have undergone certain mutations that allow them to escape these control systems. Thus, they can grow continuously to eventual form tumours. The mutations that give rise to cancer cells are diverse, and that is one of the reasons why cancer is considered a heterogeneous disease.
Breast cancer is a good example of this heterogeneity. There is a high degree of diversity between and within breast tumours, as well as among patients, and all of these factors together determine the risk of disease progression and therapeutic resistance. This implies that the cancer of each individual develops and grows in a singular way, thus causing it to be unique in terms of the treatment that it responds to. Therefore, understanding tumour cell biology is essential for improving cancer treatments.
In the lab, we assess this diversity in various ways. In this course, we will work with one the most popular approaches, namely “cell lines”, which are cells that continuously grow in vitro and resemble the characteristics of the patient they were taken from. Using a panel of cancer lines, we can screen new drugs or combinations of drugs and predict which of them will be more effective and especially the kind of patient that will benefit most from them.
5. Rational drug design
Sanja Zivanovic (Orozco’s Lab)
The typical drug discovery and development cycle, from concept to market, takes approximately 14 years and can cost up to 1 billion euros. This process could be vastly improved by combining scientific knowledge and powerful computational methods. Such an approach aims to identify the damaged protein that causes illness and also find hits and possible inhibitors, thus reducing the labour costs and time- consuming process of drug discovery.
Students will be introduced to fundamentals of molecular modeling where they will practice how to draw 2D and 3D structures. They will learn how to manipulate atoms and molecules, set up correct charge and analyze optical activity. We will go through various molecular visualization softwares: Avogadro, VMD and Pymol and carry on different graphical representation of molecules. To be able to use above mentioned softwares, students will be provided with necessary computational tools such as command line in Linux. The acquired skills in this tutorial will help them to carry out more drug discovery calculations in the second semester.
6. The value of autophagy for mitochondria
Yuliana Enciso (Zorzano’s Lab)
Autophagy (“self-eating”) is a homeostatic process by which cellular components are encapsulated into organelles called autophagosomes and then brought to the lysosome in order to be broken down and recycled for other uses. The main role of this process is the clearance of abnormal intracellular proteins and organelles.
The imbalance of this degradation pathway is involved in some pathophysiological processes, such as cancer, heart disease, and infectious disease, and in neurodegenerative conditions like Huntington’s, Alzheimer’s and Parkinson’s disease.
Most neurodegenerative diseases are characterised by the formation of protein inclusions. Therefore, understanding the proteins involved in these pathways will provide answers as to how to treat and perhaps prevent the diseases.
During this course, we will look into non-selective and selective forms of autophagy. We will then focus on a specific process that involves the degradation or removal of mitochondria. Moreover, we will explore two proteins implicated in a sporadic form of Parkinson’s disease. In the practical session, we will use cells to study the effect of some agents on mitochondria.
SEMESTER 2
1. Regulation of gene expression or why we don't understand our genome (yet)
Clara Suñer (Mendez’ lab)
In the year 2000 the Human Genome Project was completed at a cost of around 3 billion dollars and ten years of intense work. The main conclusion was that over 98% of our genetic material was junk DNA. Far from resolving our questions, reading our genome made us recognize we didn't understand it. Since then, mounting evidence has made us realize that to understand our genome means to understand how its expression is regulated.
Now we are starting to get a glimpse of how the cells counts on extremely complex regulatory networks to switch genes on and off through a myriad of novel players such as microRNAs, long non-codingRNAs and RNA-binding proteins.
Understanding how gene expression is controlled will be vital to know how an erythrocyte and a neuron are so different while sharing the exact same genes or why and individual gets a disease like cancer while and identical twin remains completely healthy.
In this course we will take a look at the current methods that researchers use to study the regulation of gene expression using in vitro cell cultured systems.
2. An introduction to neurodegeneration and neurodegenerative diseases
Elzbieta Maria Szulc (Salvatella’s Lab)
Alzheimer’s disease is the most common form of dementia. It has no cure, and symptoms worsen as the disease progresses, eventually leading to death. It was first described by German psychiatrist and neuropathologist Alois Alzheimer in 1906. The causes and progression of Alzheimer's disease are not well understood. Research indicates that this condition is associated with plaques and tangles of amyloid-beta peptide in the brain. Current treatments alleviate the symptoms of the disease, but none stop or reverse its progression. Since 2012, more than 1000 clinical trials have been conducted to find ways to treat the condition, but whether any of the treatments being tested will be effective remains to be seen. While mental stimulation, exercise, and a balanced diet have been proposed as ways to delay the onset of cognitive symptoms (though not brain pathology) in healthy elderly individuals, there is no conclusive evidence supporting the effect of these approaches. In this course, we will take an in-depth look at the mechanisms of neurodegeneration, with a focus on Alzheimer’s disease and Parkinson’s disease.
We will begin with an introduction to the history of these diseases, looking at when they were first detected in the brains of patients. This will be followed by an explanation of the molecular and cellular mechanisms involved. We will then go through the hypotheses of the causes, and the latest research strategies to find cures. Finally, we will finish the course with a question and answer session in which students will have the opportunity to reflect on future approaches in this field of research.
3. Stepping into the world of the cell skeleton (This course is a continuation from semester I)
Artur Ezquerra (Lüder’s Lab)
Microtubules are highly regulated structures involved in several cellular functions, such as the transport of proteins and organelle, migration, signalling and chromosome segregation during cell division.
Due the wide range of microtubule functions, considerable research effort is devoted to studying the mechanism that regulates the behaviour of these structures. Defects in the microtubule cytoskeleton have been linked to several human diseases, such neuropathologies and cancer.
During cell division, the microtubule network re-organises to form the mitotic spindle, a macromolecular machine that segregates the replicated chromosomes into two daughter cells. Affection in the proper organisation of the mitotic spindle leads to abnormal cell division, a critical step in cancer development.
Another field of interest in microtubule research is the cilium, a microtubule-based structure found in most mammalian cell types. Alterations in the function of cilia cause a variety of human disorders globally known as ciliopathies, some of which lead to impaired brain development and kidney diseases.
In addition, microtubules are essential for neuronal migration, polarity, and differentiation. Defective microtubule regulation in neuronal cells causes some cases of Parkinson´s and Alzheimer diseases.
For all these reasons, a better understanding of the microtubule machinery will help us develop new insights into the way to deal with many human diseases.
During the second part of the practical course, we will focus on the study of microtubule defects. Using various treatments in cultured cell lines and immunofluorescence techniques, we will identify the phases of mitosis and the effects of such treatments on cell cycle by affecting the microtubule formation.
4. Stepping into the world of proteins
Luis Daniel Toribio Jorge Garcia (Coll's Lab)
Proteins form the very basis of life. They regulate a variety of activities in all known organisms, from replication of the genetic code to transporting oxygen, and are generally responsible for regulating the cellular machinery and determining the phenotype of an organism. Proteins accomplish their tasks in the body by three-dimensional interactions between various substrates. The functional properties of proteins depend upon their three-dimensional structure. The (3D) structures arise because particular sequences of amino acids in a polypeptide chain fold to generate, from linear chains, compact domains with specific structures. The folded domains either serve as modules for larger assemblies or they provide specific catalytic or binding sites.
Different technologies are used for discovering the structure of proteins, such as X-ray crystallography. To perform this technique, we need to grow crystals, a material whose constituents, in this case proteins, are arranged in an ordered pattern that extend in all three spatial dimensions. When applied to them, crystals scatter X-rays, the information obtained from the crystals can help infer the protein structure.
5. Rational drug design (This course is a continuation from semester I)
Antonija Kuzmanic (Orozco’s Lab)
The capacity of X-ray crystallography to determine protein structures has provided scientists with an innovative way to find new drugs. By combining this approach with emerging computational methods, rational drug design not only has significantly reduced the costs of this process, but has made the treatment of many diseases possible, even on a personalized level.
This part of the course seeks to teach students how to combine structural information with computational methods, such as molecular docking, in order to discover a novel drug. Students will be given a list of proteins that are successful examples of rational drug design and will be asked to choose one that interests them. We will then study the basics of the protein structure, the features of its binding pockets, and how the possible ligands for these pockets are selected, and we will use docking software to evaluate whether the proposed compounds will make effective drugs.
6. Why mitochondrial dynamics are essential for life
Paula Martínez-Cristóbal (Zorzano’s Lab)
Mitochondria are mobile organelles present in dynamic networks. They continuously join by fusion and divide by fission. The secret of mitochondrial dynamics is revealed by confocal live-cell imaging with the use of potentiometric dyes or mitochondria-targeted fluorescent proteins.
Fission and fusion regulate fundamental cellular processes, such as calcium homeostasis and the generation of ATP and reactive oxygen species, and consequently play key roles in cell-cycle progression, apoptosis, mitophagy, and oxygen sensing. However, the disruption of mitochondrial dynamics contributes to the pathogenesis of complex diseases, including cancer, cardiovascular disease, and neurodegenerative diseases, which are not considered to involve mitochondria.
In this course, we will explore the relationship between mitochondrial dynamics, apoptosis, and autophagy. We will perform experiments with cells and work on elucidating the role of some mitochondrial proteins actively involved in mitochondrial fusion.
Venue
CRAZY ABOUT BIOMEDICINE se realizará en las instalaciones de IRB Barcelona.
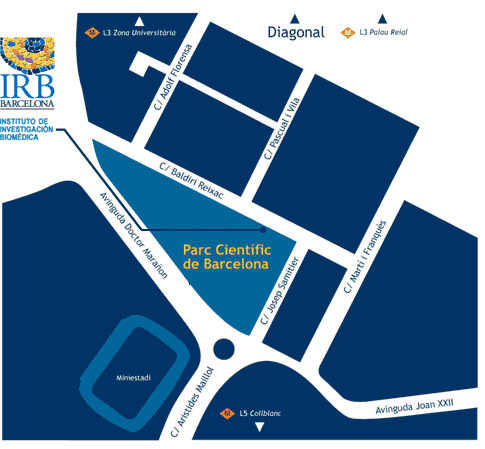
Lloc de celebració
IRB Barcelona
c/o Parc Cientific de Barcelona
Carrer Baldiri Reixac, 10
08028 Barcelona
(Campus de la Diagonal, Universitat de Barcelona)

