Mass Spectrometry & Proteomics
The Mass Spectrometry & Proteomics Core Facility (MSPCF) was created in 2007 and is currently a leading lab in the field of mass spectrometry and proteomics. The Facility team members carry extensive experience and a high level of excellence in proteomics and our lab is in continuous development and implementation of state-of-the-art methods and technologies in response to user needs. We offer a professional and personalized proteomics service to the IRB Barcelona scientific community and to public or private spheres.
For more information, please contact the facility manager (marta.vilaseca@irbbarcelona.org)

The Facility offers complete proteomics service; consultancy and experimental design, sample prep, data acquisition, data analysis, statistics and report with final discussion.
List of dedicated MS-based proteomic analysis:
- High Throughput proteomics. Untargeted approaches
- Proteome analysis:
- Label-free proteome quantitation. DDA, DIA
- Label-based proteome quantitation. TMT
- Phosphoproteome analysis:
- Label-free phospho proteome quantitation. DDA, DIA
- Label-based phosphoproteome quantitation. TMT, SILAC
- Targeted proteomics: PRM quantitative strategies
- Protein characterization. Bottom-up, top-down approaches, intact mass analysis
- PTMs analysis
- Phosphorylation differential analysis
- Denatured intact mass analysis
- Structural proteomics & protein-protein interactions
- Crosslinking-MS
- Native Mass Spectrometry. Protein complexes
- Differential interactomics
- Bio-ID-Mass Spectrometry/ Turbo-ID MS
- Chemoproteomics MS experiments
- Biofluid analysis. Differential proteomic analysis of clinical samples
- Plasma
- CFS
- Synovial Fluid
- Small molecule analysis: exact mass of purified compounds
- Launching in September: Spatial proteomic analysis
- MALDI-Imaging analysis
- High sensitivity proteomics: low input microdissected tissues/single cell
The Facility constantly works in method development and implementation to increase its competitiveness and thus to be able to provide users the most efficient solutions for their experiments with MS-based proteomic tools, in order to answer concrete biomedical questions.
- Support in experimental design
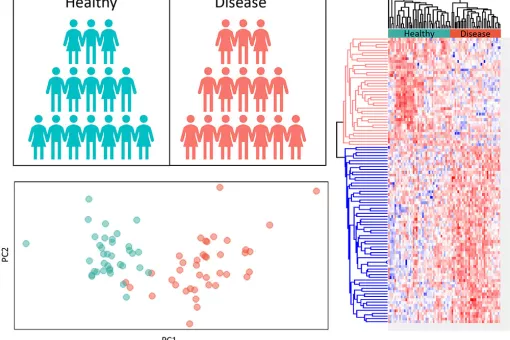
- Data analysis, statistics of all services and applications
- Functional analysis
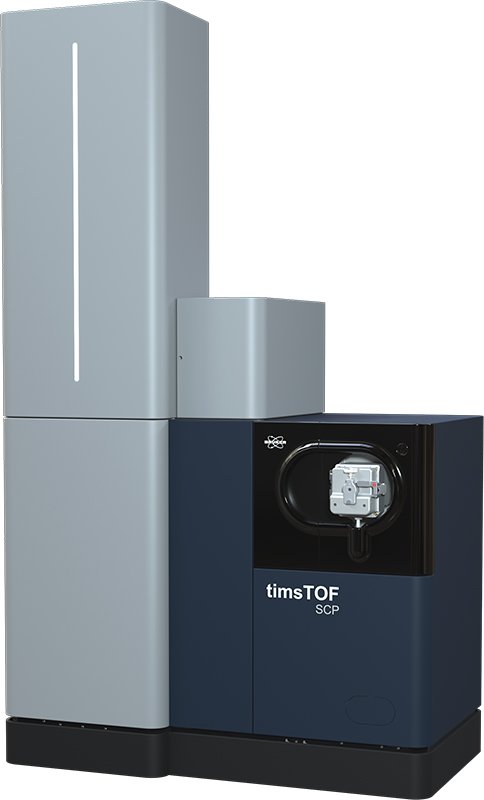
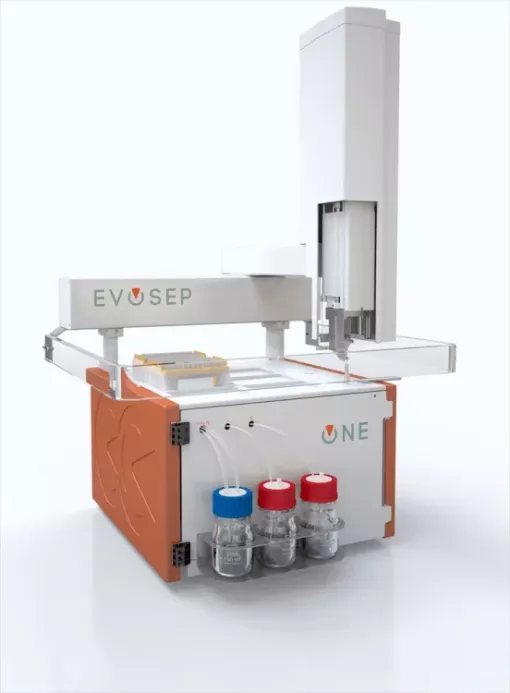
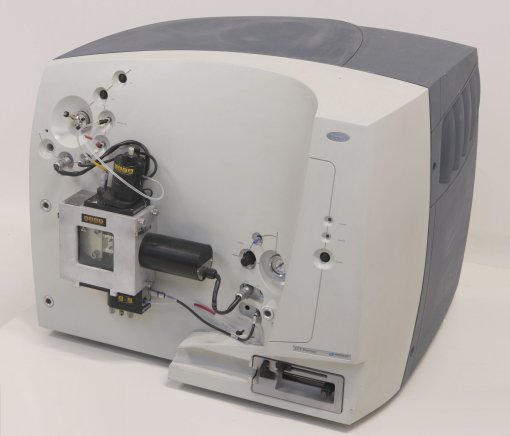





- Initial inquiry: send an email to the core facility manager marta.vilaseca@irbbarcelona.org to arrange a meeting to discuss the best approaches to reach your research goals.
- Request submission: once the project has been discussed, fill out this form
- Review & approval: CF staff reviews the request and, if needed, contacts the user for clarification. A quote is sent to the user/PI.
- Sample preparation: one of the most important steps to achieve a successful result in mass spectrometry analysis is a well-planned experiment involving adequate sample preparation. Here you have some general instructions, but it is strongly recommended to contact the MS team before starting it.
- Sample Submission: There are two ways to submit samples for MS analysis:
- Submission to the technician-run service by filling in the sample/s request form (available here or in the MSC lab).
- Self-service (open-access) systems. Only for researchers qualified and trained to operate the mass spectrometers.
If you plan to use the MS service often and would like to be considered for training, please contact marta.vilaseca@irbbarcelona.org.
When filling the request form, your sample or set of samples will be identified by a code number (apart from the users label). Depending on the analysis requested it will be classified in the corresponding "to-run" folder for each instrument.
The sample will be also classified and kept in the conditions specified in the request form.
If preferred you can keep your sample and bring it a day before the beginning of analysis (following scheduled dates).
- Service processing: samples are analysed either directly by Facility staff or by researchers (previously trained by facility members), who can use mass spectrometers through an open-access system.
- Data Analysis (if applicable): data is processed, analyzed, and interpreted according to the service or core facility scope.
- Results Delivery: results performed by MS staff will be given in printed format (spectra and report specifying analytical conditions). Digital format data can also be kept if requested. If further processing of your results is required a computer will be reserved in the facility for this purpose. Our future goal is that users can also access data via a secured network. Data will be transferred to a common-server and back-ups will be performed automatically by the ITS Department.
- Follow-up: optional meeting to explain results or assist with interpretation.
The Facility currently leads the set-up of the Biomedical Proteomics Platform (IRB Barcelona’s Unit) for biomedical and translational Research usage in Catalonia, which is co-financed by the European Regional Development Fund (ERDF) in a join collaborative project with the Josep Carreras Leukemia Research Institute (FIJC), the Sant Joan de Déu Foundation (FSJD), the Vall Hebron Institute of Oncology (VHIO) and the University of Barcelona (UB) in the framework of the 2014-2020 ERDF Operational Progamme in Catalonia (Reference: IU16-015983).
The specialization and achievements of the Facility with respect to the implementation and development of Quantitative MS and Structural Proteomics workfows within the Spanish Proteomics community has led to its partnership in the PRBB network (Plataforma en red de Recursos Bioinformaticos y Biomoleculares, former ProteoRed), obtaining financial support from the Spanish Government-Instituto de Salud Carlos III for this purpose in the periods 2014-2017 and 2018-2020 (PRB3 (IPT17/0019 - ISCIII-SGEFI / ERDF), as a member group of Proteored, PRB3-ISCIII). The Facility participates in the international Chromosome-Centric Human Proteome Project through the Spanish HPP consortium (SpHPP), In this regard, the Facility is contributing Top-down data to the “Chromosome 16 study”,
During the period 2014-2018, the IRB Barcelona Facility was granted a COST Action (European Cooperation in the field of Scientific and Technical Research-COST) from the Horizon 2020 program in a collaborative consortium initially formed by 8 European countries. The action was entitled “Native Mass Spectrometry and Related Methods for Structural Biology” (Action BM1403), aiming to bring together a group of researchers with a common interest, namely the development and application of new biomolecular MS methods in order to make the characterization of protein structure and dynamics more rapid and routine.
There is a calendar (contact the facility manager) for each instrument and submissions will be ordered by time of entry (IRB Barcelona groups are given preferential treatment).
Given the diversity of applications, some of which require only fast analysis while others require longer analysis time, the calendar will be organised as follows:
- Users can reserve a week in a specific instrument for long application (maximum 2 full weeks per user, depending on demand).
- Between long time applications we always reserve a week to perform shorter analysis in order to avoid delays.
- Mondays (or a pre-specified day) are reserved for the maintenance of the equipment and set up of the required technique pretended to use that week in each instrument.
Data will be kept for a maximum of two years.