Microtubule organization in cell proliferation and differentiation

Meet Our Scientists Videos
Research information
Background
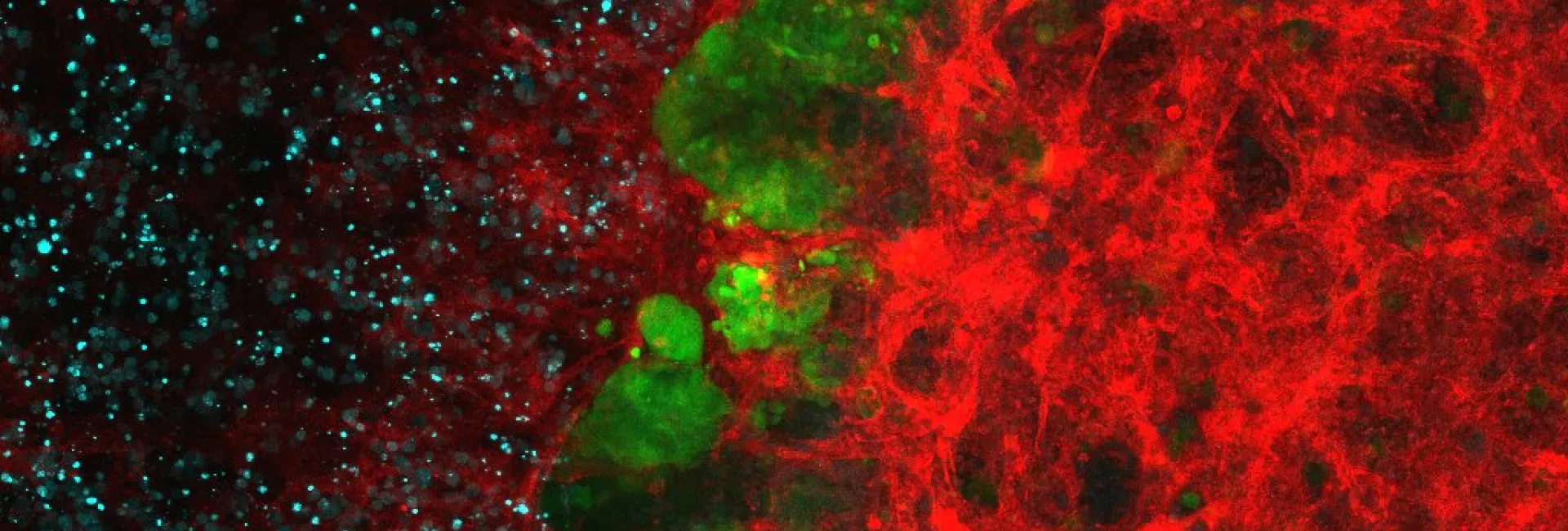
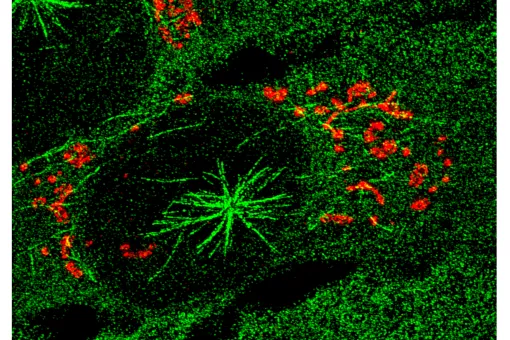
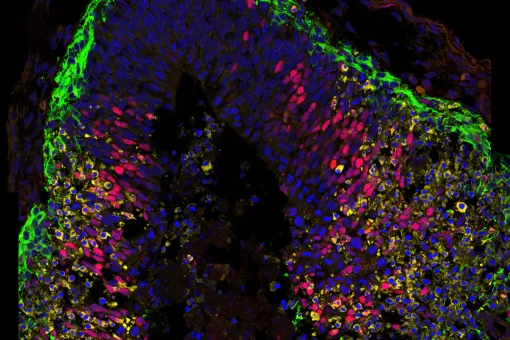
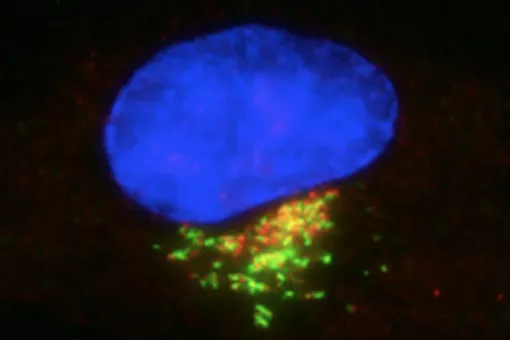
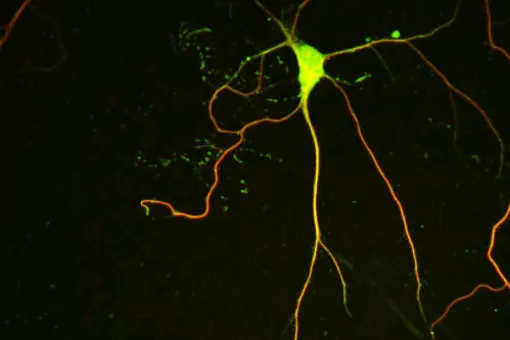
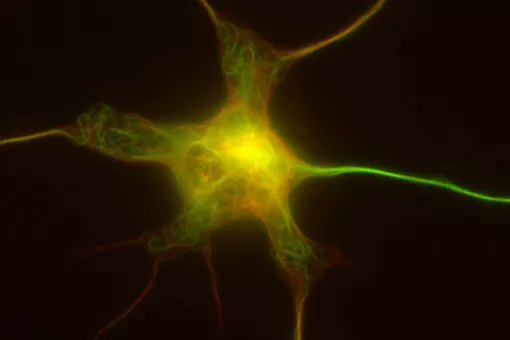
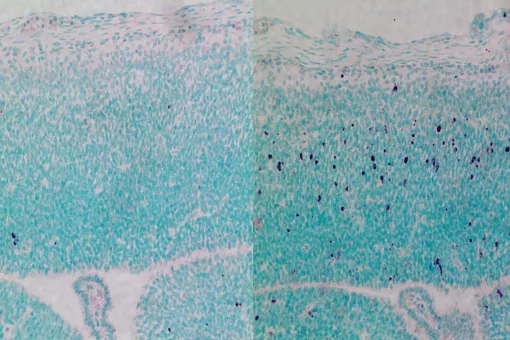
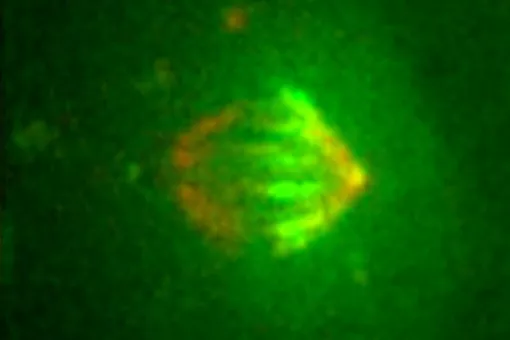
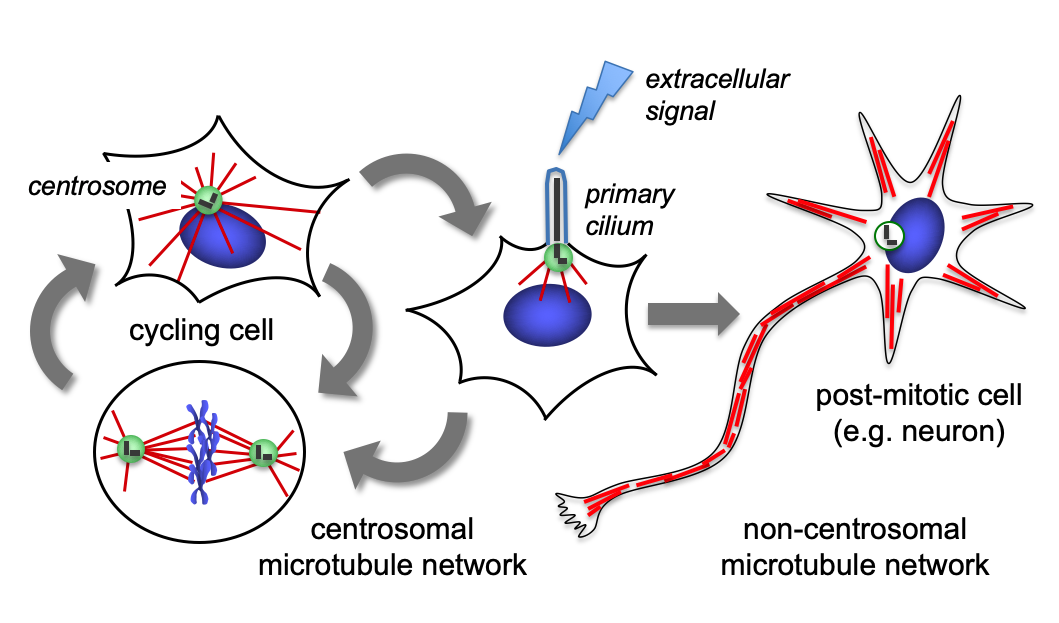
Microtubules are filamentous polymers composed of α- and β-tubulin and form part of the cytoskeleton. Microtubules organize the cell’s interior and mediate essential processes such as intracellular transport or the segregation of chromosomes during cell division. To carry out these functions, microtubules need to be arranged as ordered arrays with specific polarity and distribution. Two mechanisms are key to achieving this: spatial and temporal control over the formation of new microtubules, a process termed nucleation, and the attachment of microtubules to cellular structures collectively known as microtubule organizing centers (MTOCs). Centrioles are particularly important for the microtubule cytoskeleton, since they do not only function as MTOC (known as centrosome) but are also required for the assembly of cilia. Cilia are elongated, membrane-enclosed structures that project from the cell surface. They can be motile and used, for example, for clearing mucus from the surface of epithelial cells in the trachea of our lungs, or immotile, functioning as sensory organelles that receive and relay extracellular signals to the nucleus, to regulate gene expression. Ciliary signaling is crucial for cell fate decisions that govern proliferation and differentiation. Mutations in genes encoding proteins that constitute or regulate the function of microtubules, MTOCs or cilia can have severe consequences and have been linked to a range of diseases including neurodevelopmental and neurodegenerative disorders as well as cancer.
Research interests
Microtubule-dependent processes such as segregation of chromosomes during cell division require microtubules to be organized as highly ordered arrays. Failure to properly organize microtubules impairs tissue formation and function, causing developmental disorders, and can contribute to cancer development.
The overall goal of our lab is a molecular understanding of how cells generate and remodel microtubule arrays with specific configurations during cell cycle progression and during cell differentiation, and how errors are linked to disease. We also seek to unravel the biogenesis and function of centrosomes and cilia, two key structures of the microtubule cytoskeleton.
We study these structures and processes in cultured human cells including induced pluripotent stem cells (iPSCs) and iPSC-derived tissues, in combination with functional assays after knockdown or CRISPR/Cas9-mediated genome editing, and high-resolution microscopic imaging.
In addition, for a better molecular and mechanistic understanding, we aim to reconstitute key components and processes in vitro, using recombinant protein expression and purification.
Research lines
How are centrosomes and cilia assembled and how do they function?
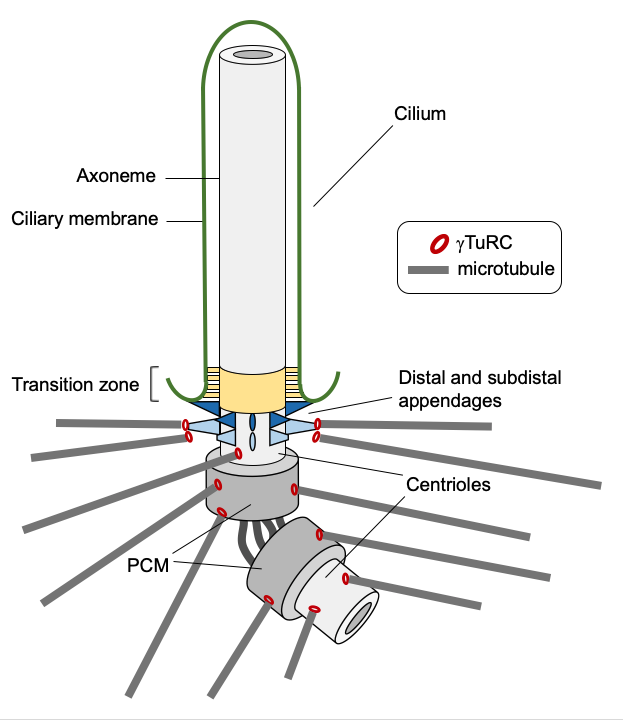
The centrosome is composed of a pair of centrioles, surrounded by a proteinaceous pericentriolar material (PCM). The PCM harbors the microtubule nucleating activity, which at least in part depends on the γ-tubulin ring complex (γTuRC) and is crucial for the centrosome to function as MTOC. We aim to understand how γTuRC is recruited to centrioles and how recruitment is coupled to γTuRC activation, to convert centrioles into MTOCs.
Apart from functioning as MTOCs, centrioles are essential for the formation and function of cilia. We are interested in how MTOC function is coordinated with cilium assembly and disassembly both during the cell cycle and during cell differentiation.
How is the microtubule cytoskeleton organized in non-transformed cells and how is it remodeled during their differentiation?
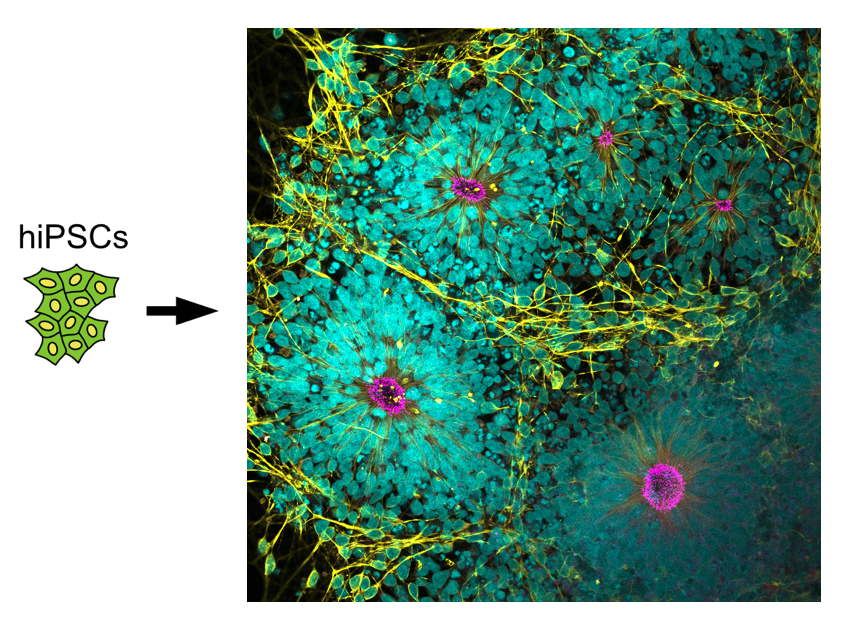
Most of our molecular understanding regarding the structures and mechanisms that constitute and organize the microtubule cytoskeleton was obtained through functional studies in transformed cancer cell lines. However, recent work suggests that non-transformed cells implicated in normal development as well as numerous diseases may employ different or additional, yet-to-be-discovered mechanisms to organize their microtubule networks. To close this knowledge gap, we characterize the roles of known MTOCs such as the centrosome and of novel, non-centrosomal MTOCs in human iPSCs and derived cell types during tissue formation.
How does the microtubule nucleator γ-tubulin ring complex (γTuRC) nucleate microtubules and how is it regulated?
An important question is how cells control the formation of new microtubules, which is crucial for assembly and remodeling of microtubule arrays. Microtubule formation does not occur spontaneously, but requires a nucleator such as γTuRC. γTuRC was proposed to provide a template for the assembly of new microtubules. A major recent advance contributed by our lab was the in vitro reconstitution of human γTuRC, which now allows a detailed structure-function analysis of the nucleation mechanism and of its regulation. Cryo-EM analyses of purified γTuRC have revealed a surprising asymmetry that is incompatible with the template nucleation model. How γTuRC gets activated to become an efficient nucleation template is a major open question.

How do defects in the microtubule cytoskeleton cause disease?
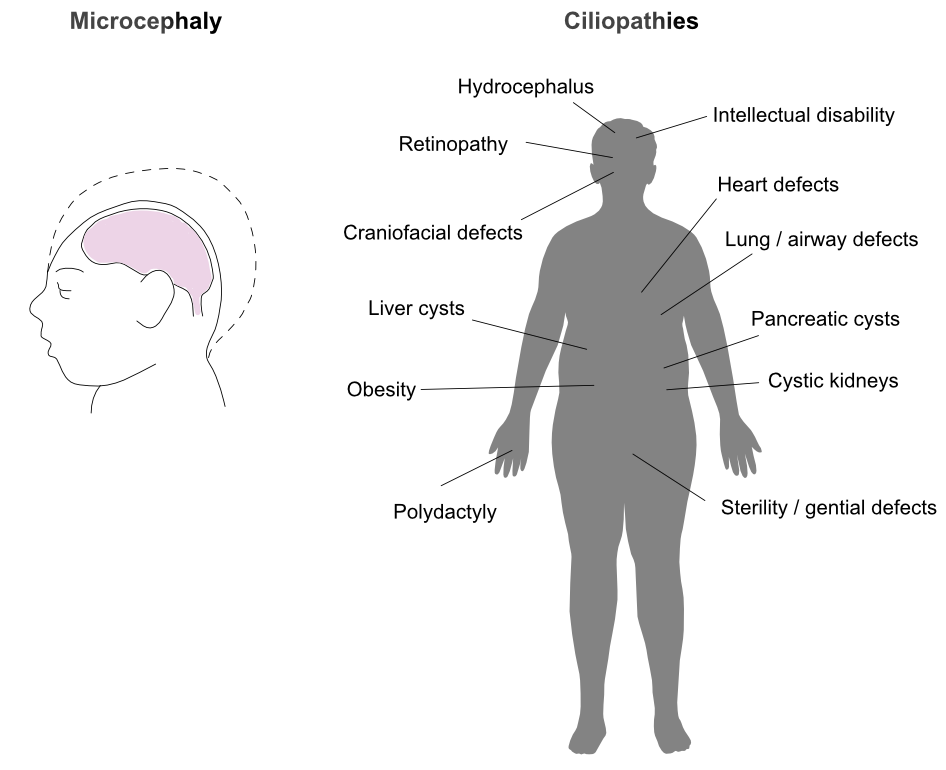
Mutations in genes encoding proteins that constitute or regulate centrosomes are associated with impaired developmental growth, in particular during brain development, resulting in microcephaly (reduced brain size). Defective formation or function of cilia has been linked to disorders collectively known as ciliopathies, which affect the development and function of a wide range of body parts and organs. We aim to use insight generated in our analyses of fundamental, microtubule cytoskeleton-related processes to unravel the molecular and cellular causes of associated diseases.
A recent example is Cohen Syndrome, a rare developmental disorder, characterized by a range of clinical features including developmental delay, microcephaly, intellectual disability, obesity, low muscle tone, neutropenia (a reduction in immune cells that fight infections), and progressive retinopathy (loss of vision). Unfortunately, due to limited awareness and funding opportunities, many rare diseases including Cohen Syndrome are understudied and there are no or only limited treatment options. Thus, basic research elucidating the underlying molecular and cellular causes is urgently needed. Since we suspect an involvement of the microtubule cytoskeleton in Cohen Syndrome, we leverage our expertise in this area and work with patient families (through the Spanish Cohen Syndrome Association SICOES) (https://www.facebook.com/profile.php?id=100076486994683), and treating clinicians, to establish cell and tissue models to study the disease.
If you are interested in fundraising or donating for our research, please get in touch. Additional information can also be found here (https://www.irbbarcelona.org/en/donate/testimonials).
Selected publications
Projects
"Dissecting gamma-TuRC composition and activity by Single Molecule Pull-down" (GTR), financiado mediante el Programa de Innovación e Investigación de la Unión Europea Horizonte 2020 bajo la acción Maire Sklodowska-Curie (IF). Referencia: 703907.
“Linking cellular defects with clinical manifestations in Cohen Syndrome”, financiado por La Marató de TV3. Referencia: 202019-30

"Remodelación de los microtúbulos del citoesqueleto y destino celular durante el desarrollo cerebral" (REMOBRAIN), cofinanciado por el Ministerio de Ciencia, Innovación y Universidades- Agencia Estatal de Investigacióny por el Fondo Europeo de Desarrollo Regional (FEDER) de la Unión Europea. Referencia: PGC2018-099562-B-I00



Grup de Recerca consolidat (SGR 2017-2019) de la Secretaria d'Universitats i Recerca del Departament d'Empresa i Coneixement de la Generalitat de Catalunya. Agencia de Gestió d'Ajuts Universitaris i de Recerca (AGAUR). Referencia: 2017 SGR 1089.


"Exploring the molecular mechanisms that link microtubule organization to brain development and neuroregeneration" (MONDAR), cofinanciado por el Ministerio de Economía y Competitividad y por el Fondo Europeo de Desarrollo Regional (FEDER) de la Unión Europea. Referencia: BFU2015-69275-P.


“A new concept to explain the regulation of microtubule dynamics" (MICRODYN), financiado por el Ministerio de Economía, Industria y Competitividad - Agencia Estatal de Investigación (Convocatoria Explora Ciencia 2015). Referencia: BFU2015-72951-EXP.


“Explorant noves estratègies per tractar la neutropènia en pacients amb síndrome de Cohen” Agència de Gestió d’Ajuts Universitaris i de Recerca. Referència: 2021 LLAV 00025

“Membrane homeostasis as potential therapeutic angle in Cohen Syndrome” Orphan Disease Center & University of Pennsylvania. Reference: MDBR-22-136-Cohen


“Rescue of defective lipid handling in Cohen syndrome” Orphan Disease Center & University of Pennsylvania. Reference: MDBR-24-17-Cohen


“Organización de los microtúbulos y su rol en la formación de tejidos” Ministerio de Ciencia, Innovación y Universidades- Agencia Estatal de Investigación y Fondo Europeo de Desarrollo Regional (FEDER) de la Unión Europea. Referencia: PID2021-127603NB-I00

“Modulación de la nucleación y anclaje de microtúbulos en distintos tipos celulares” Ministerio de Ciencia, Innovación y Universidades- Agencia Estatal de Investigación. Referencia: PID2024-156734NB-I00

Policies
Candidates who would like to join our group should e-mail a cover letter explaining their motivation and interest in our work, a CV, and contact information for referees to jens.luders![]() irbbarcelona.org.
irbbarcelona.org.