Images
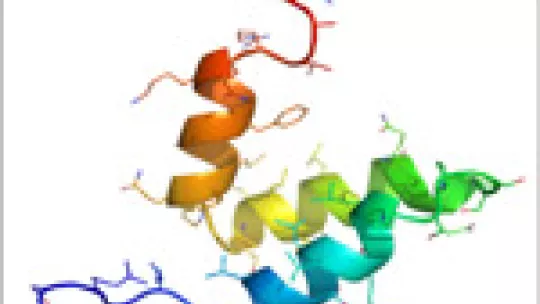
Proteins, the molecular machines that carry out the majority of the functions within a cell, are typically seen as possessing a specific three-dimensional structure. Cellular processes (e.g. reproduction) are activated or deactivated by a molecular choreography that involves the binding of one protein to another. Of late, it has been recognized that even a small protein (say, of size 50 residues) can binding to multiple partners – this results in an advantageous situation for the cell wherein depending on the identity of the partner a different functional response can be generated. In parallel, it has become increasingly evident that a single static structure is a poor representation of a protein and that it is more apt to represent them as a collection of conformations (the ‘ensemble’). This leads to the realization that the motion of different protein parts (the ‘dynamics’) is critical for function. The question that has therefore challenged scientists is how does a small protein show such promiscuity and what is the underlying molecular basis.
To approach this problem, Athi N. Naganathan (Juan de la Cierva fellowship) and Modesto Orozco from the Joint IRB Barcelona-BSC program in Computational Biology chose the nuclear co-activator binding domain (NCBD) of the CREB DNA-binding protein (CBP), a small a-helical domain of around 50 residues, as a test case example. Despite its size, NCBD displays remarkable promiscuity – it binds to seven different partners. They have performed extensive molecular dynamics simulations of NCBD both from reduced and all-atom representations, and as constraints employ a variety of experimental techniques that report on the dimensions and degree of structure in the ensemble. They reconstruct for the first time an atomic level representation of the functional ensemble of NCBD and show that multiple conformations co-exist thus answering fundamental questions on the functional binding mechanism. Simulations show that NCBD displays the characteristics of a downhill folder – i.e. the transition from completely unfolded to the folded structure occurs without encountering any significant free-energy barrier. They report on how the downhill folding mechanism of NCBD is intricately tied to its functional requirement thus highlighting a unique mechanism (the ‘molecular rheostat’) to achieve multi-functionality in small binding domains.
Reference articleThe Native Ensemble and Folding of a Protein Molten-Globule: Functional Consequence of Downhill Folding.
Athi N. Naganathan and Modesto Orozco.
JACS (dx.doi.org/10.1021/ja204053n)
About IRB Barcelona
The Institute for Research in Biomedicine (IRB Barcelona) pursues a society free of disease. To this end, it conducts multidisciplinary research of excellence to cure cancer and other diseases linked to ageing. It establishes technology transfer agreements with the pharmaceutical industry and major hospitals to bring research results closer to society, and organises a range of science outreach activities to engage the public in an open dialogue. IRB Barcelona is an international centre that hosts 400 researchers and more than 30 nationalities. Recognised as a Severo Ochoa Centre of Excellence since 2011, IRB Barcelona is a CERCA centre and member of the Barcelona Institute of Science and Technology (BIST).