Images
A study published in Nature Communications and led by Maria J Macias highlight new DNA motifs for the Smads proteins.
The main TGF-β signal transduction mechanism is the Smad pathway. As transcription factors, Smad proteins interact directly with DNA. However, the motif/s specifically recognized by Smads have been a matter of debate for many years. In the past, an in vitro approach identified the 5’–GTCT–3’ but its 4-base short length suggested that its presence in many Smad target genes could be random. Recent experiments in live cells have revealed that TGF-β target genes bind Smad2/3 and Smad4 complexes in promoters and enhancers lacking GTCT motifs.
A study headed by ICREA researcher Maria J Macias at the Institute for Research in Biomedicine (IRB Barcelona) has discovered a new high affinity 5-nucleotide consensus that demonstrates how Smad4, the conserved component in Smad complexes, and also Smad3 bind to these sites, providing new grounds for the understanding of cytokine-driven transcriptional control. The results indicate that the concept of Smad Binding Element needs to be revised in the literature and in textbooks, to include the new findings. The study has been published in Nature Communications in December.
Identification of the 5GC GGC(GC)|(CG) site
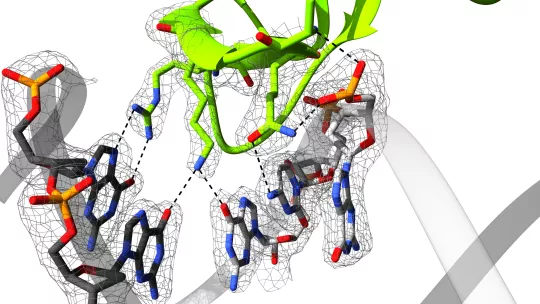
Using a combination of experimental techniques, the researchers identified a consensus GC-motif as the binding site of Smad4 and Smad3, and they provided 2.05 Å resolution X-ray crystal structures of the protein-DNA complexes. “Using CRISPR/Cas9, we demonstrated that binding of Smads to these sites is functional,” says Maria J. Macias. Remarkably, Smad3 and Smad4 bind to the GGC(GC)|(CG) sites reading up to five bases. The flexibility of the DNA binding hairpin characterized by NMR and SAXS, facilitates access to DNA duplexes with slightly distinct topologies and DNA sequences.
The scientists also found that the new 5GC-motifs are highly represented as clusters in Smad-bound regions genome-wide. Binding to clusters suggest how three MH1 domains in a trimeric Smad complex may bind to promoters/enhancers at once with high specificity. This capacity endows Smad transcriptional complexes with a level of adaptability that has established TGF-β as one of the most versatile and highly conserved signaling pathways in metazoans. The findings also highlight regions in promoters affected by tumour mutations, which have not been regarded as Smad targets so far.
This study was done in collaboration with the labs headed by Joan Massagué at the Memorial Sloan Kettering Cancer Center, in New York, and José A. Márquez at the European Molecular Biology Laboratory, in Grenoble.
The study was funded by the Ministry of Economy and Competitiveness, the BBVA Foundation and NIH grants. Access to the HTX facility at the EMBL (Grenoble) was supported by the Horizon 2020 Programme grant iNEXT of the European Commission.
Article of reference:
Pau Martin-Malpartida, Marta Batet, Zuzanna Kaczmarska, Regina Freier, Tiago Gomes, Eric Aragón, Yilong Zou, Qiong Wang, Qiaoran Xi, Lidia Ruiz, Angela Vea, José A. Márquez, Joan Massagué & Maria J. Macias
Structural Basis for Genome Wide Recognition of 5-bp GC Motifs by SMAD Transcription Factors
Nature Communications (2017) doi:10.1038/s41467-017-02054-6
About IRB Barcelona
The Institute for Research in Biomedicine (IRB Barcelona) pursues a society free of disease. To this end, it conducts multidisciplinary research of excellence to cure cancer and other diseases linked to ageing. It establishes technology transfer agreements with the pharmaceutical industry and major hospitals to bring research results closer to society, and organises a range of science outreach activities to engage the public in an open dialogue. IRB Barcelona is an international centre that hosts 400 researchers and more than 30 nationalities. Recognised as a Severo Ochoa Centre of Excellence since 2011, IRB Barcelona is a CERCA centre and member of the Barcelona Institute of Science and Technology (BIST).