Images
Of the 614 cell lines studied, 7% closely corresponded to a different type of cancer than that thought to give rise to the cell line.
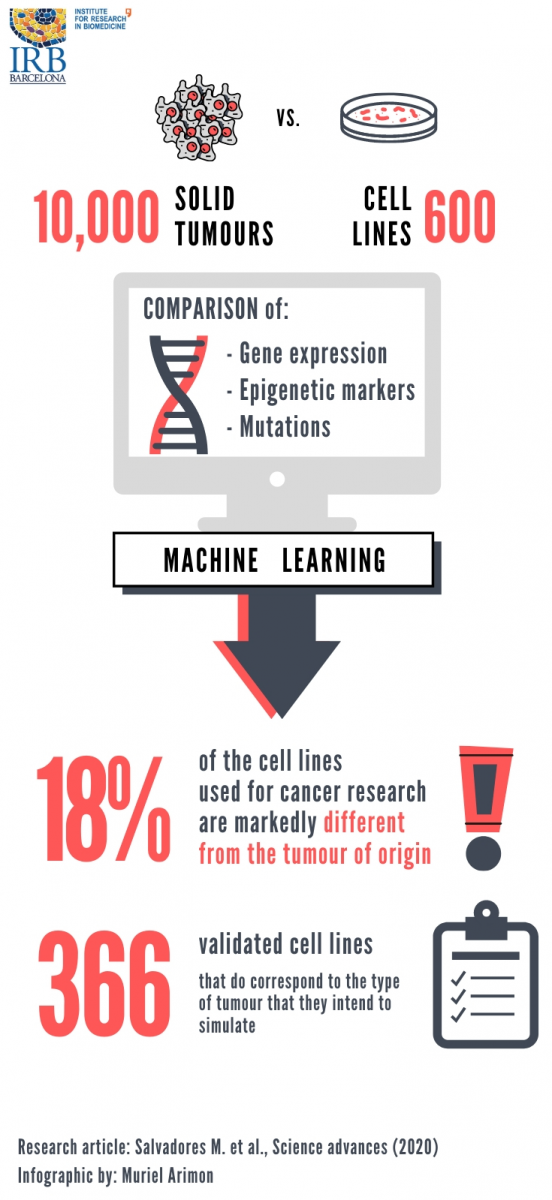
The Genome Data Science Laboratory at IRB Barcelona has drawn up a reference list of 366 cell lines in which the genetic pattern corresponds to the type of tumour that they intend to model.
The work shows that studies using the cell lines on this reference list have a greater discovery rate, and has been published in the journal Science Advances.
In cancer research, cell lines are an essential tool to study tumour biology and to test drugs or therapeutic approaches. Led by the ICREA researcher Fran Supek, the Genome Data Science Laboratory at IRB Barcelona has evaluated the adequacy of 614 cell lines and has concluded that only approx. 60% greatly conserve the characteristics of the tumour of origin, while another 20% maintain some resemblance. The remaining 18% differ considerably from the tumour that they supposedly represent. Therefore, they provide a less reliable model of the tumour type they are meant to represent. Surprisingly, 7% of all cancer cell lines used in labs worldwide closely correspond to a completely different type of cancer than that they are originally believed to be.
Different types of cancer, such as colon or lung cancer, are distinct diseases that evolve and are treated in a different manner. Cell lines for cancer research are typically generated from a surgical tissue sample from a patient's tumour. They are grown in the laboratory and serve as a study model for that disease. For research projects that target a particular type of cancer, it is crucial that the cells used in the laboratory truly represent the type of cancer they are supposed to represent.
 “Identifying cell lines that are very different from the tumour of origin is crucial because they can lead to false positive or false negative results in experiments,” says Supek. "Promising new candidates for cancer drugs may remain undiscovered because the tissue-of-origin of cell lines was not correctly identified," he explains.
“Identifying cell lines that are very different from the tumour of origin is crucial because they can lead to false positive or false negative results in experiments,” says Supek. "Promising new candidates for cancer drugs may remain undiscovered because the tissue-of-origin of cell lines was not correctly identified," he explains.
Large-scale data analysis with immediate application
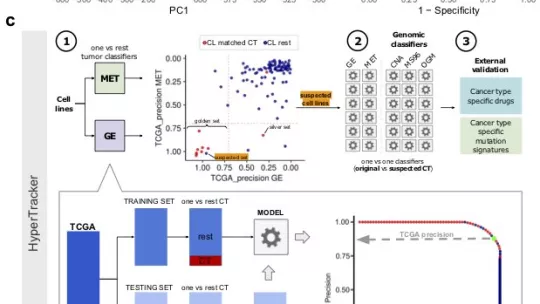
This is the first large-scale data analysis that jointly considers genetic, gene expression and epigenetic patterns of more than 600 cancer cell lines, together with 9,000 samples from 22 types of human tumours. Using machine-learning methods, the Genome Data Science Laboratory has compiled a list of 366 cell lines that reliably correspond to the tumour of origin. This list is available as a resource to the scientific community as a part of the article, published in the journal Science Advances.
The researchers have also analysed the results of previous experiments by other labs that have used many cell lines in parallel. They have shown that, by eliminating misclassified cell lines or using only cell lines from the reference list, we are better placed to discover new drugs that target specific markers. Their data analyses point to several new molecules that may show activity against pancreatic or brain cancer cells that bear particular mutations.
“We hope that prioritizing the cell lines that we have selected will help researchers obtain more reliable experimental results when searching for new cancer treatments,” says Supek.
Ovarian cancer or skin cancer?
One of the cell lines studied, ES-2, derives from sample of an ovarian tumour surgically removed from a 47-year-old woman. However, the scientists observed that the pattern of activated and silenced genes shown by this cell corresponds to that of skin cancer. "Then we studied the specific mutations in this cell line, and we saw that they have a mutational signature that is caused by UV rays. We think that this ovarian tumour may have in fact originated from a metastasis of a skin cancer and that is why it has these characteristics,” says Marina Salvadores, the lead author of this study. “In total, we identified five different cell lines that may originate from skin cancer rather than the various internal organs that they were thought to come from. This seems consistent with the high propensity of melanoma to metastasize to various organs.”
Marina Salvadores is a PhD student with the Genome Data Science Laboratory of the IRB Barcelona and she holds a FPU fellowship. Fran Supek is an ICREA research professor, head of the Genome Data Science Laboratory and an EMBO Young Investigator. The work in the laboratory is funded by the ERC Starting Grant “HYPER-INSIGHT” and the Spanish Ministerio de Ciencia e Innovación grant “RegioMut”.
Reference article:
Marina Salvadores, Francisco Fuster-Tormo & Fran Supek
Matching cell lines with cancer type and subtype of origin via mutational, epigenomic, and transcriptomic patterns
Science Advances (2020) DOI:10.1126/sciadv.aba1862
About IRB Barcelona
The Institute for Research in Biomedicine (IRB Barcelona) pursues a society free of disease. To this end, it conducts multidisciplinary research of excellence to cure cancer and other diseases linked to ageing. It establishes technology transfer agreements with the pharmaceutical industry and major hospitals to bring research results closer to society, and organises a range of science outreach activities to engage the public in an open dialogue. IRB Barcelona is an international centre that hosts 400 researchers and more than 30 nationalities. Recognised as a Severo Ochoa Centre of Excellence since 2011, IRB Barcelona is a CERCA centre and member of the Barcelona Institute of Science and Technology (BIST).