Images
An international team led by computational biologist Fran Supek at IRB Barcelona develop a machine learning method to predict unknown gene functions of microbes.
The system examines and compares ‘big data’ available on the metagenomes of human and environmental microbiomes.
Understanding the functions of genes in bacteria that form part of the human microbiome—the collection of microbes found inside our bodies—is important because these genes might explain mechanisms of bacterial infection or cohabitation in the host, antibiotic resistance, or the many effects—positive and negative—that the microbiome has on human health.
Surprisingly, the functions of a huge number of microbial genes are still unknown. This knowledge gap can be thought of as “genomic dark matter” in microbes, and neither computational biology nor current lab techniques have been able address this gap.
This challenge has now been tackled through an international collaboration between the Institute for Research in Biomedicine (IRB Barcelona) and two other interdisciplinary research centres, namely the IJS in Ljubljana (Slovenia) and RBI in Zagreb (Croatia). The findings have been published recently in Microbiome, the international journal of reference in microbiome research. The study was led by Fran Supek, computational biologist and leader of the Genome Data Science lab at IRB Barcelona, and first-authored by Vedrana Vidulin, a computer scientist affiliated to the centres in Slovenia and Croatia.
Intelligent prediction method
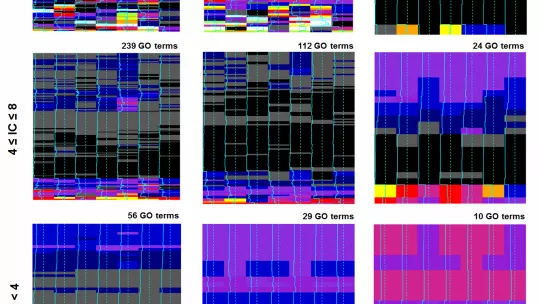
The researchers have developed a new computational method able to examine thousands of metagenomes simultaneously and identify the evolutionary signal that can predict the function of many microbial genes. This method, which analyses “big data” from human microbiomes (e.g. from the intestine or skin) and other metagenomes (e.g. from the soil or ocean) is based on a special kind of machine learning algorithm: it can create "decision trees” to predict hundreds of different functions at once, finding links between genes and at the same time predicting what they do in the microbial cell.
“This makes the algorithm very good at not getting confused by the noise in the metagenomic data, meaning that it is accurate and can confidently propose a biological role for a large number of genes with unknown functions. Intriguingly, it also proposes many additional functions for genes that already have some known role,” says Supek.
The most important finding to emerge from this research is that the analysis of human microbiomes and other metagenomic data, such as those of the soil and ocean, allows researchers to assign hundreds of gene functions that have evaded current computational genomics approaches until now. “In other words, metagenomes allow scientists to see what ordinary genomes don’t,” explains the Croatian researcher, who was recently awarded a grant from the European Research Council (ERC).
Diversity is key
The scientists have found that different types of environments can predict different types of gene functions. For example, metagenomes from the ocean can be used to predict the genes used by bacteria for photosynthesis. But as the researchers point out, this could not have been discovered from the bacteria in the human gut. In contrast, the gut microbiome has been very useful for predicting key genes involved in the mechanisms underlying the development of disease and in the metabolism of alcohol and the biosynthesis of certain amino acids—predictions that would have been more difficult to make using microbiomes from the environment.
The authors conclude that, through machine learning, a large and diverse set of environments allows us to learn about many different gene functions in microbes. “Computational methods like this one are shedding light on the “dark matter” within microbial genomes –- the enormous number of genes in bacteria and in archaea whose functions are a mystery,” says Supek.
The thousands of computational predictions generated will need to be validated in experiments. Once validated, they may lead to the discovery of new genes that explain how bacteria shape the ecosystems around us and indeed the ecosystem within us—the human microbiome.
This study has been funded through the European FP7 ‘Future and Emerging Technologies’ Programme and an ERC Starting Grant.
Reference article:
The evolutionary signal in metagenome phyletic profiles predicts many gene functions
Vedrana Vidulin, Tomislav Šmuc, Sašo Džeroski and Fran Supek
Microbiome (2018) 6:129 Doi: https://doi.org/10.1186/s40168-018-0506-4
VÍDEO MEET OUR SCIENTISTS. Fran Supek: "Solving the riddle of DNA"
About IRB Barcelona
The Institute for Research in Biomedicine (IRB Barcelona) pursues a society free of disease. To this end, it conducts multidisciplinary research of excellence to cure cancer and other diseases linked to ageing. It establishes technology transfer agreements with the pharmaceutical industry and major hospitals to bring research results closer to society, and organises a range of science outreach activities to engage the public in an open dialogue. IRB Barcelona is an international centre that hosts 400 researchers and more than 30 nationalities. Recognised as a Severo Ochoa Centre of Excellence since 2011, IRB Barcelona is a CERCA centre and member of the Barcelona Institute of Science and Technology (BIST).