Images
A team of British and Spanish researchers have developed COCO, a computer tool to increase the possibility of modelling proteins using data derived from nuclear magnetic resonance. The study, which is published the April issue of the journal Proteins, seeks to contribute to the information held in protein data banks.
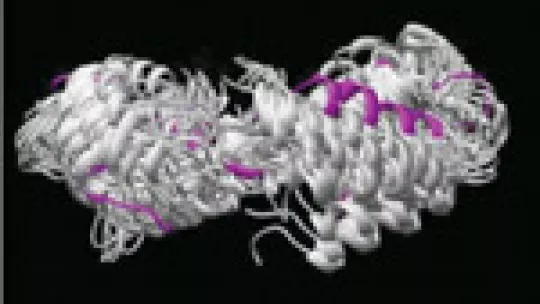
“Nuclear magnetic resonance (NMR) experiments provide, among others, data on protein conformation, and COCO (from English, COmplementary COordinates) is a server developed to explore whether any protein conformation coherent with these data is overlooked”, explains Modesto Orozco, at the Institute for Research in Biomedicine (IRB Barcelona).
The researchers use data from NMR technology and propose “ensembles of protein structures” that best fit the same. This information is deposited in international databases, such as the PDB (Protein Data Bank), but does not reflect the structural diversity allowed by NMR data.
The server COCO, whose features are described in this month’s issue of the journal Proteins, analyses the distribution of the proposed ensembles of structures and generates another ensemble that offers new solutions that better satisfy the data. This computer tool is based on principal component analysis, a statistical technique used to integrate and process information in data banks.
According to its creators, the programme is “very fast” as it can analyse ensembles of up to 25 elements in two minutes using a conventional computer. The software is freely accessible on the website of this international research project in computational biology (http://www.ccpb.ac.uk/).
COCO is the result of collaboration between the University of Nottingham and the European Bioinformatics Institute, both in the UK, with the Spanish institutions IRB Barcelona, the “Instituto Nacional de Bioinformática” and the Barcelona Supercomputing Center (BSC).
Related article:
“COCO: a simple tool to enrich the representation of conformational variability in NMR structures”.
Laughton C.A., Orozco M., Vranken W.
Proteins 75(1): 206-216, Abril de 2009.
About IRB Barcelona
The Institute for Research in Biomedicine (IRB Barcelona) pursues a society free of disease. To this end, it conducts multidisciplinary research of excellence to cure cancer and other diseases linked to ageing. It establishes technology transfer agreements with the pharmaceutical industry and major hospitals to bring research results closer to society, and organises a range of science outreach activities to engage the public in an open dialogue. IRB Barcelona is an international centre that hosts 400 researchers and more than 30 nationalities. Recognised as a Severo Ochoa Centre of Excellence since 2011, IRB Barcelona is a CERCA centre and member of the Barcelona Institute of Science and Technology (BIST).