Images
This species shows several genetic duplications, which may explain its high plasticity and capacity to adapt to diverse farming conditions.
CSIC has headed this research, which has brought about a considerable advance in the development of genomic and biotechnological tools to achieve more sustainable fish farming.
The findings of the study will be applied in genetic selection and environmental programming in order to improve fish quality and interaction with the microorganisms present in the host environment.
The gilthead seabream is of great economic importance, being widely farmed throughout the Mediterranean and reaching an annual production of more than 200,000 tonnes. Its success in aquaculture is attributed to its hermaphrodite nature, with a male-female switch after the second year of life, as well as its capacity to adapt to changes in salinity, temperature, oxygen availability and food composition. To unravel the genetic basis of the plasticity of this species, researchers belonging to the Nutrigenomics and Fish Pathlogy groups at CISC´s Institute of Aquaculture Torre de la Sal (IATS-CSIC), in collaboration with the company Biotechvana S. L. and the Center for Genomic Regulation (CRG) in Barcelona, have headed a four-year study devoted to sequencing its genome.
The reason why this species has many genes and gene duplications
A massive genomic sequencing and assembly strategy has allowed the scientists to gain the most complete vision of the genomic sequence of this species to date. “We have sequenced and assembled more than 1,250 million base pairs, out of an estimated total of 1,600 million,” explains Jaume Pérez-Sánchez, research professor at CSIC and leader of the project. The genome of this species is considerably larger than that of other species of interest in fish farming such as sea bass and turbot. Consequently, more than 55,000 genes that are actively expressed have been identified, although the most remarkable finding is that more than half of these, specifically 55%, have multiple copies. “Many of these are generated by mobile genetic elements,” says Carlos Llorens, CEO of Biotechvana.
“A large number of these multiplications do not correspond to evolutionary duplication events already known in fish, but are instead typical of this species and therefore recent in evolutionary terms,” says Toni Gabaldón, group leader at CRG. In addition, the scientists have demonstrated than many of the duplicated genes are involved in generating a response to environmental changes, such as immune or sensory responses, as well as in genomic transcription. Therefore, it is believed that the duplication of the molecular machinery confers this species with capacity to adapt to domestication and to hostile environments or scenarios of global change—“this capacity requiring rapid and appropriate responses to intensive farming systems that have a high pathogen load,” comments Ariadna Sitjà-Bobadilla, a member of the Fish Pathology Group at IATS.
Examples of the phenotypic plasticity of the seabream
The gilthead seabream, therefore, provides an excellent model in which to study the processes that lead to genomic expansion in higher vertebrates and the effects of this process on the phenotypic plasticity of the species. As proof of this, fattening protocols performed at IATS farming facilities in Castellón demonstrate that the metabolism of a carnivore like the gilthead seabream is sufficiently adaptable to achieve 1-kg specimens with only 1.2 Kg of plant-based feed with a low content of fish flour and fish oil, according to the dietary formulations proposed in the European project ARRAINA.
Likewise, work carried out in collaboration with researchers at the Universidad de las Palmas de Gran Canaria, within the framework of the European project PerformFish, and published recently in the journal Epigenetics (doi: 10.1080/15592294.2019.1699982) has demonstrated that “it is possible to nutritionally program progeny by making changes to parental diets that affect key genes in lipid metabolism, such as delta 9 desaturase,” according to Jaume Pérez-Sánchez.
Other examples of the phenotypic plasticity of the gilthead seabream have been evidenced in the framework of the European project Infrastructures in Aquaculture, AQUAEXCEL2020. In this regard, Josep Calduch-Giner, member of the Nutrigenomics Group at IATS, says, “by subjecting animals to low concentrations of oxygen early on, we can metabolically reprogram them so that months later they are able to maximise energy production aerobically when challenged with low oxygen conditions, for example during the period of fast growth that occurs in the summer months.” “We do not know the mechanisms responsible for this, but changes observed in the intestinal microbiota may play a key role,” says Carla Piazzon, member of the Fish Pathology Group at IATS.
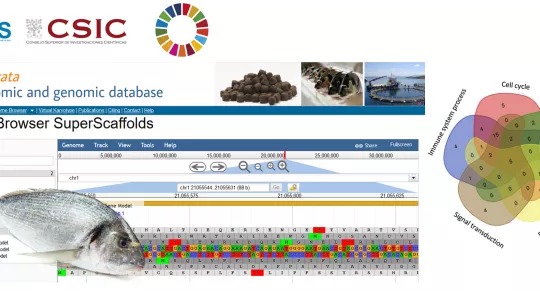
The researchers have placed a genome viewer of the gilthead seabream on their website, thus making it available to the public and scientific community (www.nutrigroup-iats.org/seabreamdb). The research has received national (Proyecto Intramural del CSIC 1201530E025; BreamAquaINTECH, RTI2018-094128-B-I00) and European (proyecto AQUAEXCEL2020) funding and has been published in the journal Frontiers in Marine Science.
Jaume Pérez-Sánchez, Fernando Naya-Català, Beatriz Soriano, M. Carla Piazzon, Ahmed Hafez, Toni Gabaldón, Carlos Llorens, Ariadna Sitjà-Bobadilla and Josep A. Calduch-Giner. Genome sequencing and transcriptome analysis reveal recent species-specific gene duplications in the plastic gilthead sea bream (Sparus aurata). Frontiers in Marine Science 6:760. DOI: https://doi.org/10.3389/fmars.2019.00760
About IRB Barcelona
The Institute for Research in Biomedicine (IRB Barcelona) pursues a society free of disease. To this end, it conducts multidisciplinary research of excellence to cure cancer and other diseases linked to ageing. It establishes technology transfer agreements with the pharmaceutical industry and major hospitals to bring research results closer to society, and organises a range of science outreach activities to engage the public in an open dialogue. IRB Barcelona is an international centre that hosts 400 researchers and more than 30 nationalities. Recognised as a Severo Ochoa Centre of Excellence since 2011, IRB Barcelona is a CERCA centre and member of the Barcelona Institute of Science and Technology (BIST).