ICREA Research Professor
Meet Our Scientists Videos
Research information
Background
Our research addresses the evolution of the protein synthesis machinery, the molecular interactions that regulate it, and the biomedical applications that can be derived from its study. Our research projects are focused around the biology of transfer RNA (tRNA). This adaptor molecule is responsible for the faithful transmission of genetic information to protein sequence. tRNAs are ancient macromolecules that witnessed the origin of life on earth, and today act as universal regulators of gene translation.
We are interested in the mechanisms of tRNA aminoacylation in mitochondria, and how this reaction is used to coordinate the cycle of mitochondria with the rest of the cell’s metabolism. One of the goals of this research is to increase our understanding of human mitochondrial diseases caused by translation defects in this organelle.
We also investigate the biological role of adenosine deaminases of tRNAs (ADATs). ADATs are essential enzymes that drove the evolution of the eukaryotic genome. Their biochemical properties are relatively well understood, but the mechanisms behind their biological relevance remain unclear. We are working towards the identification of specific human genetic programs regulated by ADAT.
Research interests
First, we strive to clarify the mechanisms that integrate the process of protein synthesis with the rest of the cell’s biology. We do that by investigating factors that control mitochondrial translation, and by studying enzymes that catalyze tRNA modifications.
In parallel, using tRNA, aminoacyl-tRNA synthetases, and tRNA modification enzymes as tools, we want to improve our understanding of the evolution of life in general, and of the eukaryotic cell in particular.
Finally, we are intent on converting our academic understanding of tRNA biology into biomedical applications. This approach is justified by the large body of experimental data that has accumulated on tRNA aminoacylation, and by the essential role of this reaction in biology.
Research lines
a. Mitochondrial aminoacylation, metabolic control, and disease models.
We are currently characterizing the aminoacylation activity of the mitochondrial enzyme seryl-tRNA synthetase (SRS). Animals use two forms of this enzyme, one in the cytosol and one in mitochondria. The misacylation or poor activity of mitochondrial SRS in humans is responsible for a variety of metabolic diseases. However, these conditions are often due to mutations in the mitochondrial genome, which is extremely difficult to reproduce and study. We have characterized mitochondrial SRS and, through its mutation, we have generated a Drosophila model that can be used as a proxy for human mitochondrial disease in a model system that is easy to manipulate.
In the process of investigating mitochondrial SRS in Drosophila we discovered a new essential mitochondrial protein which we named SLIMP. SLIMP is an important regulator of mitochondrial translation that possibly controls the replication status of mitochondria and somehow ties this process to the cell cycle. We are currently investigating the molecular mechanisms behind this coordinating role of SLIMP.
b. The importance of aminoacylation editing in human cells.
Central to maintaining a healthy proteome is the interaction between codon and anticodon, which links the genetic information in the form of triplets in the messenger RNA to the amino acid brought in by the transfer RNA. This interaction takes place at the core of the ribosome, and ensures that the protein that is being synthesized incorporates the right amino acid in the right place and at the right speed to ensure its proper folding and activity.
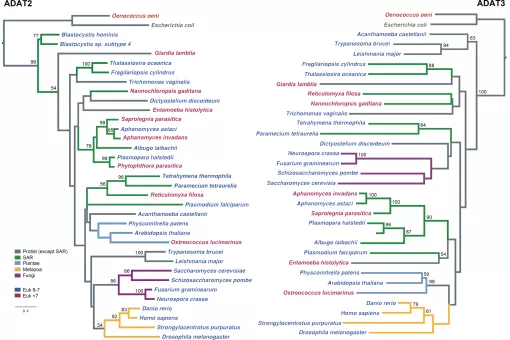
We have recently demonstrated that, in eukaryotes, this interaction is greatly influenced by an enzyme named adenosine deaminase acting on tRNAs (ADATs). ADATs constitute a family of tRNA modification enzymes related to the more general group of RNA adenosine deaminases. The function of all these enzymes is the deamination of adenosine bases to generate inosine, a simpler nucleotide with a wider pairing ability than adenine.
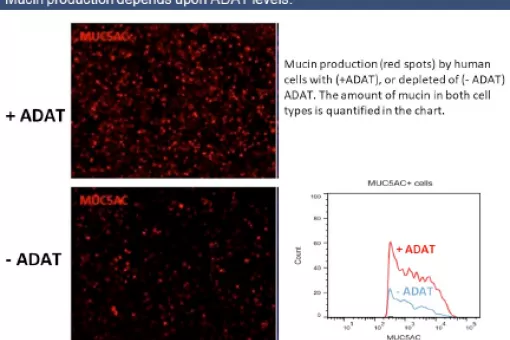
In our group we investigate the physiological importance of ADAT in animal and cellular models. This is the first functional characterization of ADAT in pluricellular eukaryotes, and our initial results show that the activity of the enzyme is absolutely essential in both organisms. The suppression of ADAT expression in whole adult flies or in muscle is completely lethal, despite the fact that the maximum levels of repression achieved are of 30% of the wild-type amounts.
In parallel to our animal studies, and to further explore the connection between ADAT function and proteome composition, we are characterizing the human genome to identify those genes that are most influenced by the activity of ADAT.
Selected publications
Projects
Grup de Recerca consolidat "Biologia de la Traducció Genetica" funded by the Agencia de Gestió d'Ajuts Universitaris i de Recerca (AGAUR) 2014 SGR 771

"Límites funcionales del código genético" funded by Ministerio de Economia y Competitividad BIO2014-61411-EXP

"Aplicaciones biomédicas derivadas del estudio de la traducción génica" funded by Ministerio de Ciencia e Innovación - Agencia Estatal de Investigación and Fondo Europeo de Desarrollo Regional (FEDER) de la Unión Europea. BIO2015-64572-R

"Engineered mutagenic TRNAS for the treatment of pancreatic cancer" funded by Fundación Científica Asociación Española contra el Cancer (AECC) IDEAS211022RIBA

"Función, regulación, y aplicaciones biotecnológicas de la síntesis de proteínas" funded by Ministerio de Ciencia e Innovación - Agencia Estatal de Investigación PID2019-