Organitzat per l'IRB Barcelona en col·laboració amb la Fundació Catalunya La Pedrera.
Els tutors d'aquesta edició són: Panagiotis Giannios, Enric Ros, Marta Frigolé, Josep Biayna and Marina Villamor, Jordi Badia, Lorena González, Joel Paz Domínguez, Manuel Cañete, Miguel Rovira, Daniela Romão, Alícia Llorente, Sara Gregorio, Oriol Pich i Inés Sentís.
Presentation

Objectius
Bojos per la Biomedicina és un curs dirigit als estudiants del primer any de batxillerat que desitgin explorar alguns dels descobriments fascinants que s'estan fent actualment en les ciències de la vida. A través d'aquest curs, els estudiants tindran l'oportunitat d'aprofundir el seu coneixement de la teoria i tècniques científiques en el camp de la biomedicina. Treballaran juntament amb investigadors joves per experimentar com es fa ciència en un institut de recerca internacional, guanyar una mica d'experiència pràctica en les últimes metodologies d'avantguarda i posicionar-se per a una possible carrera professional en les ciències de la vida.
Descripció del curs
Taller d'un any de durada sobre les ciències de la vida per a estudiants de batxillerat. Organitzat per IRB Barcelona dins el Programa "Bojos per la Ciència" de la Fundació Catalunya La Pedrera.
Aquest curs combina sessions teòriques i activitats experimentals pràctiques, que es duran a terme durant 18 dissabtes de l'any. El curs tractarà 12 temes científics actuals, que van des de la biologia cel·lular i molecular fins a la biologia estructural i computacional i la química, presentats per estudiants de doctorat i postdocs de l'IRB Barcelona. En el primer «semestre» (gener-abril 2018), els 3 primers dissabtes es dedicaran a aquestes sessions teòriques generals per a tots els participants. Durant els 6 dissabtes següents, es formaran grups petits que entraran als laboratoris per a les sessions pràctiques. A continuació, es repetirà aquest programa amb 6 temes de recerca nous per al segon semestre (maig-novembre). Els estudiants participants s'hauran de comprometre a assistir al curs durant tot l'any.
 Idioma del curs
Idioma del curs
Totes les xerrades i sessions pràctiques es faran en anglès.
Dates i horaris
El curs tindrà lloc de gener a desembre de 2018, 10.00h-14.00h.
1r SEMESTRE
- Div. 12 gener 2018: Inauguració al Mon Sant Benet- Sant Fruitós del Bages
- Diss. 13 gener: Xerrada 1
- Diss. 20 gener: Xerrada 2
- Diss. 3 febrer: Xerrada 3
- Diss. 17 febrer: Taller 1
- Diss. 3 març: Taller 2
- Diss. 17 març: Taller 3
- Diss. 7 abril: Taller 4
- Diss. 21 abril: Taller 5
- Diss. 28 abril: Taller 6
2n SEMESTRE
- Diss. 26 maig: Xerrada 1
- Diss. 16 juny: Xerrada 2
- Diss. 30 juny: Xerrada 3
- Diss. 15 setembre: Taller 1
- Diss. 29 setembre: Taller 2
- Diss. 20 octubre: Taller 3
- Diss. 27 octubre: Taller 4
- Diss. 17 novembre: Taller 5 - MODIFICADA
- Diss. 24 novembre: Taller 6
Preu del curs
Lloc on es realitzarà el curs
Institut de Recerca Biomèdica (IRB Barcelona)
C/ Baldiri Reixac, 10
08028 Barcelona
Qui pot sol·licitar una plaça
Aquest curs està dirigit als estudiants de primer any de batxillerat que tinguin un interès i talent especials en els camps relacionats amb les ciències de la vida (principalment biologia i química).
Els estudiants poden sol·licitar la plaça a un màxim de 2 dels programes de la série "Bojos per la Ciència" i participar només en un.
Com sol·licitar una plaça

Els estudiants interessats hauran d'emplenar el formulari web de sol·licitud i incloure una carta de motivació. També es demanarà una carta de recomanació directament de dos dels seus professors que coneguin bé l'alumne/a. En el cas de que l'estudiant hagi canviat de centre aquest curs, suggerim que sol·licitin les cartes als antics professors. La data límit d'inscripció (AMPLIADA) és el 30 d'octubre 2017.
El curs està obert a un total de 24 estudiants. Se seleccionaran els candidats en funció del seu expedient acadèmic, de les recomanacions dels seus professors i de la seva motivació per participar-hi. Es convidarà els candidats preseleccionats a fer entrevistes amb els organitzadors científics al novembre, després de les quals es farà la selecció final. La primera setmana de desembre es comunicarà el resultat als estudiants. Es demanarà als estudiants seleccionats per participar-hi i als seus pares/tutors legals que signin una carta de compromís d'assistir a totes les sessions.
Col·laboradors
Facebook: @LaPedrera.Fundacio
Twiter: @PedreraFundacio
Instagram: #bojosperlaciencia
Facebook: @LaPedrera.Ciencia
Twiter: @PedreraCiencia
Instagram: @lapedrera_ciencia


Per a qualsevol dubte, siusplau contacteu amb: irb_outreach@irbbarcelona.org
Dates importants
 30 d'octubre 2017: Data límit d'inscripció AMPLIADA
30 d'octubre 2017: Data límit d'inscripció AMPLIADA- 24 d’octubre-5 de novembre 2017: Contacte amb els candidats preseleccionats
- 24 octubre-12 de novembre 2017: Entrevistes
- Setmana del 27 de novembre 2017: Contacte amb els candidats seleccionats
- 30 de novembre-1 dedesembre 2017: Publicació del llistat dels estudiants acceptats
- Div. 12 de gener 2018: Inauguració del curs al Món Sant Benet –Sant Fruitós del Bages
- Diss. 13 de gener 2018: Inici del curs
- Cerimònia de clausura - Pendent de confirmació
Programme
SEMESTER 1
1. Pluripotency control mechanisms during development
Panagiotis Giannios (Development and morphogenesis in Drosophila)

An important feature of organs, is their ability to maintain their morphology and function in spite of environmental or genetic stimuli that cause natural or accidental cell loss. This capacity is often sustained by the so-called stem cells, which are able to provide new cells of different types in each tissue.
A common characteristic shared by these pluripotent cells, is their ability to remain in a “dormant” status while being able at any given point to activate their regenerative potential. The mechanisms controlling this ability have to be precisely regulated while balancing between the need to maintain tissues’ homeostasis and having to retain the limit of uncontrolled cellular growth. In mechanistic terms, this delicate balance is sustained during development by a continuous feedback loop between metabolic shifts, response to hormonal cues and regulation of gene expression.
Drosophila melanogaster, is a model organism used extensively over the past decades in several developmental studies. The well structured knowledge we now have in several aspects of the Drosophila’ s biology, allows us to use this organism as an in vivo model in several studies, including the functional characterization and the dissection of the molecular mechanisms that govern the regulation of stem cells.
In this course, we will take advantage of the powerful Drosophila genetic tools to mark progenitor cells and modify the expression levels of selected genes that regulate some of their distinct characteristics. This will give us the opportunity to study basic mechanisms that keep these cells in a multi-potent status and identify crucial aspects of their role throughout the development.
2. The code of life
Enric Ros (Gene translation laboratory)

The information encoded in the genetic material is translated into proteins following a set of rules known as the Genetic Code, which is universal for almost all the living organisms and establishes the basis for life. This means that, from the simplest bacterium to us, we all use a highly similar “language” to synthesize all the required proteins, a language based on the combination of four nucleotides (A, U, G and C). The combination of these nucleotides in groups of three (the codons) represent the “words” that all living organisms “read” in the ribosomes to elongate a chain of amino acids, eventually generating proteins.
Malfunctions of the different biomolecules participating in this highly regulated system can lead to disease, but in recent years strategies to modify some of their functions have been applied for biotechnological purposes.
In this course we will study the role of the different key constituents of the translation machinery in cells, and the essential importance of the Genetic Code in the generation of active proteins. At the practical side, we will learn the common laboratory techniques used for protein expression and purification in a prokaryotic system (Escherichia coli), and also certain specific techniques to introduce mutations in the sequence of genes to produce recombinant proteins with modified functions.
3. Chemical synthesis of bioactive molecules
Marta Frigolé (Laboratory of Molecular Biophysics)

Chemistry is at the heart of the drug-discovery process. For each new drug reaching the market, there have been billions of compounds synthesised and tested. Why so many? Because is by chemically modulating the structure of the molecules that we can fine-tune their potency and also their pharmacodynamic/pharmacokinetic properties (absorption, distribution, metabolism, excretion and side-effects).
In this practical course we will synthesise acetaminophen, also known as paracetamol or the active ingredient in Gelocatil®. While doing so we will learn how chemists plan and design the synthesis routes as well as the main techniques and instruments used to build the active ingredients of all small-molecule drugs.
4. Meet the guardians of the DNA
Josep Biayna (Genome Data Science)
Marina Villamor (Genomic Instability and Cancer Laboratory)
![]()

Our DNA can break due to external (i.e. the sun and chemical agents) and internal (i.e. DNA replication) factors. To solve this problem, we have some genes that function as the guardians of our genome, by identifying these lesions and activating the DNA Damage Response (DDR) pathway. However, the DDR pathway does not always work properly. If the damage is severe, the cell undergo to apoptosis (programmed cell death) or senescence (permanent cell cycle arrest). On the other hand, if the stability of the genome is not massively compromised, the cell survives and starts proliferating in an uncontrolled manner. The transformation of a normal cell into a tumour cell is a multi-step process that involves dynamic changes in the genome at both the genetic and epigenetic levels. Currently, cancer accounts for 8 million deaths per year, being the most common cause of death in Europe after cardiovascular diseases.
Our lab is interested in deciphering the potential role of genomic instability and the DDR in cancer predisposition. In this practical course, we will monitor the effect of DNA damage in different cancer cell lines and perform some techniques that are daily used in the lab to study this phenomenon.
5. Targeting the Tumour Microenvironment in colorectal cancer metastasis
Jordi Badia (Colorectal cancer laboratory)

Colorectal Cancer (CRC) is the most common cancer and the second leading cause of cancer-related deaths in Spain, according to the AECC (Asociación Española Contra el Cáncer). CRC produces cancers in the large intestine through a multi-step process that, like many other types of cancer, starts with the acquisition of mutations in key genes. These mutations then go on to make cells proliferate without control. CRC can be easily treated if detected early. However, we still need to find effective treatments for the most advanced stages of cancer, particularly for metastasis—the final and deadliest step of cancer, which in the case of CRC occurs mainly in the liver.
The great potential of cancer cells resides not only in their capacity for mass proliferation but also in their ability to trick the healthy cells surrounding them. Fibroblasts, cells of the immune system, and blood vessels form what we call the tumour microenvironment (TME), which protects cancer cells from being removed and facilitates their invasion of other healthy organs. Our laboratory has recently shown that the TME is crucial for the aggressiveness of CRC and formation of metastasis in the liver. We are now channelling efforts into studying the potential of the TME as a way to block metastasis and into identifying novel treatments.
In this practical course we will provide an overview of several in vivo models and histological tools. Using these tools, we will discover and target those cells that form the TME of liver metastasis, with the aim to learn how to exploit them and stop the CRC metastasis.
6. Targeting cell signaling pathways to treat cancer
Lorena González (Signalling and Cell Cycle Laboratory)

Cells have to constantly deal with changes in their extracellular environment. It means that they have to respond efficiently to these changes in order to prevent damage but also they have to control proliferation, survival and migration. To do that, cells have developed several mechanisms that allow them to receive, integrate and interpret signals to produce the appropriate response. These mechanisms, called ‘signaling pathways’, are based in a series of chemical changes. One of the most important modifications is phosphorylation and proteins which are able to phosphorylate others are called kinases.
Our laboratory is mainly focused in a particular protein kinase called p38 MAPK, which is activated under different stress situations and plays a critical role in inflammation, cell growth, proliferation, differentiation and cell death. However, when this pathway is dysregulated diseases such as cancer can arise. In that case, p38 MAPK inhibition could be beneficial to treat patients.
In this practical course we will provide an overview of the p38 MAPK signaling pathway and which are the main techniques we use in the lab to study it. We will use purified proteins and cultured cell lines to learn how p38 MAPK inhibitors work. Finally, we will explore which patients could benefit from them.
SEMESTER 2
1. The Role of Microtubules in Cell Division and Cancer
Joel Paz Domínguez (Microtubule organization)

Cellular cytoskeleton is perhaps one of the most important structures of the cell as it is involved in several functions as morphogenesis, cellular division or cell migration. It is composed by three members, actin filaments, intermediate filaments and microtubules. Microtubules are hollow cylindric polymers composed by tubulin dimers. Microtubules are very dynamic structures that constantly undergo episodes of polymerization and depolimerization which is known as dynamic instability. Microtubules are involved in key processes as intracellular transport of cargos (proteins, vesicles and even organelles) and in mitosis where they are in charge of the formation of the mitotic spindle and proper chromosome segregation. One of the hallmarks of cancer is uncontrolled cell division and one approach to treat cancer is to try to kill cancer cells. Compounds that target microtubules disrupting the normal cell division cycle and eventually lead to cell death have been proven to be one of the most effective cancer chemotherapeutic drugs available.
In this practical course, we will learn how a mitosis occurs in vivo, what are the different phases of mitosis, what is the role of microtubules in this process and how can we treat cancer focusing on microtubules. For this purpose, we will use cultured cell lines treated with several chemotherapeutic drugs used in cancer therapy and see how they affect cell division with the help of fluorescence light microscopy techniques.
2. Regulation of gene expression or why we don't understand our genome (yet)
Manuel Cañete (Translational control of cell cycle and differentiation)

In the year 2000, the Human Genome Project was completed at a cost of around 3 billion dollars and ten years of intense work. The main conclusion was that over 98% of our genetic material was junk DNA. Far from resolving our questions, by reading our genome it became apparent that we didn't understand it. Since then, mounting evidence has made us realise that understanding our genome means understanding how its expression is regulated.
We are now starting to gain some insight into how cells depend on extremely complex regulatory networks to switch genes on and off through a myriad of novel players such as microRNAs, long non-coding RNAs, and RNA-binding proteins. Unravelling the mechanisms underlying the regulation of gene expression will be vital to understand how an erythrocyte and a neuron differ while sharing the exact same genes or why in identical twins one falls ill with a disease like cancer while the other remains completely healthy.
In this course we will use in vitro cell culture systems and look at the current methods that researchers apply to study the regulation of gene expression.
3. Strategies to understand ageing and senescence
Miguel Rovira (Cellular Plasticity and Disease)

Ageing is characterised by a gradual deterioration of physiological function, and it is considered the primary risk for human pathologies such as cancer or cardiovascular disorders. One of the mail hallmarks of ageing is cellular senescence, the phenomenon by which normal cells cease to divide. Senescent cells accumulate in the organism over time, secreting pro-inflammatory molecules and contributing to ageing-related diseases.
There is an increasing interest in clinical medicine to better identify and target senescent cells, as their elimination delays and ameliorates some ageing-associated diseases, and can even extend longevity.
Students will acquire hands-on experience using different techniques to induce cellular senescence in normal cells. They will learn state of the art techniques to specifically target senescent cells, analysing in vitro and in vivo samples. Finally, they will also have the opportunity to perform classical protocols to detect senescent cells such as the SAβgal staining. These techniques will help us to understand, identify and target senescent cells, which is a promising therapeutic approach in clinical medicine.
4. Drosophila as a model organism for studying cancer
Daniela Romão (Development and Growth Control Laboratory)

Cancer is driven by complex genetic and cellular mechanisms that ultimately cause abnormal cells to undergo uncontrolled proliferation (tumor formation) and invasion of healthy tissues (metastasis).
The most common type of human tumor is the carcinoma. Carcinoma is derived from epithelial tissue, such as the skin, but it can become invasive or metastatic by spreading beyond the primary tissue layer to surrounding tissues or organs. These transition is mediated by the activation, in aggressive cancer cells, of the EMT programme (Epithelial to Mesenchymal Transition) causing them to undergo morphogenetic alterations that allows a higher invasion capacity.
The fruit fly, Drosophila melanogaster, with a long and impressive history as a model organism for studying epithelial development, revealed itself as a potent tool for studying carcinomas offering the possibility of analyzing the detailed interaction between cells, tissues and genes. Due to the fact that this fly maintains conserved the signaling pathways present in humans, we also have the possibility of reducing genome redundancy when establishing comparations between the two.
In this practical course, we will learn about Drosophila has a model organism for studying cancer related mechanisms. We will identify different fly structures such as the wing imaginal disc and the fat body through in vivo dissections and perform immunohistochemistry in order to understand how different signaling pathways are analyzed.
5. In vivo experimental approach to study Breast Cancer Bone Metastasis
Alicia Llorente and Sara Gregorio (Growth control and cancer metastasis)


In a primary tumor, many cells are constantly trying to enter into the circulation, travel through the bloodstream and arrive to a new organ to form a secondary tumor. This process is called metastasis and is very inefficient. However, tumor cells sometimes acquire properties that give them the ability to succeed and form a metastasis, which usually is fatal for cancer patients.
Metastasis is a very complex process, that is why we need different tools and techniques for its study. Despite in vitro experimentation gives us the possibility to study different aspects of tumor cells behavior inside the lab,in vivoexperimentation allows a deeper understanding of how this tumor cells develop in their natural environment.
In our lab we combine all these techniques in order to study breast cancer metastasis. We use human cancer cell lines and animal models to unravel the role of specific genes in breast cancer bone metastasis.
In this course we will learn how cells are cultured and labeled in the lab, the techniques that we use to inject them into mice, how tumor growth can be monitored and the correct way to analyze the results obtained using in vivo models.
6. Uncovering the genomic makeup of thousands of tumors: implications for cancer personalized medicine
Oriol Pich and Inés Sentís (Biomedical Genomics)


Cancers is a disease where cells start to grow and proliferate in an uncontrolled way. These cells accumulate somewhere from a few to millions of changes in their DNA, which are called somatic mutations. But only a subset of them confer malignant properties (driver mutations) whereas most have little or no impact on how the cell functions (passenger mutations). The finding of driver mutations is essential for understanding the biology of cancer and developing the so-called precision medicine, which relies on identifying the driver mutations of the patient to prescribe drugs specifically targeting them. We call this process insilico drug prescription.
Analyzing thousands of tumors with all their mutations is a challenging task –luckily we can use computers to help us analyze and interpret this vast amount of biological data (bioinformatics!). During the program the students will be introduced to the fundamentals of Bioinformatics and Cancer Genomics. They will use and write several programs in the Python programming language to analyze the mutations of thousands of tumors from real patients, and perform a state-of-the-art insilico drug prescription according to the driver mutations found. They will also be introduced to the most common forms of visualization of cancer mutational data. No prior programming skills are required.
Venue
El programa BOJOS PER LA BIOMEDICINA es durà a terme a les instal·lacions de l'IRB Barcelona.
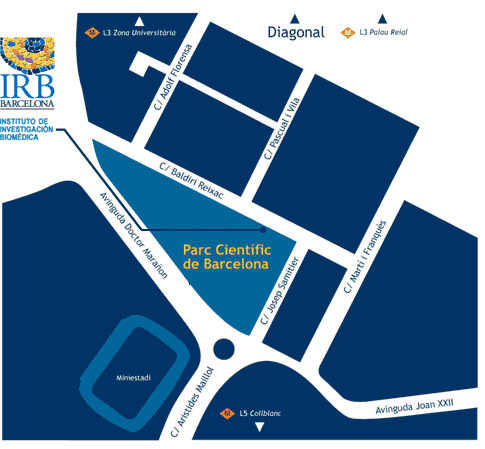
Institut de Recerca Biomèdica (IRB Barcelona)
Parc Cientific de Barcelona
C/ Baldiri Reixac, 10
08028 Barcelona
(Campus de la Diagonal, Universitat de Barcelona)

