Images
E. coli is the best known model organism at the molecular level. It is used extensively for biotechnology production, and its infectious strains are of biomedical interest.
The article in Nature Biotecnology is a collaborative effort between biologists, biochemists and bioinformaticians at the Craig Venter Institute, the University of Virginia, and the Institute for Research in Biomedicine (IRB).
The team of 18 researchers presents the first map of key interactions between the molecular components of the bacteria, and allow us to understand how its most intimate machinery works.
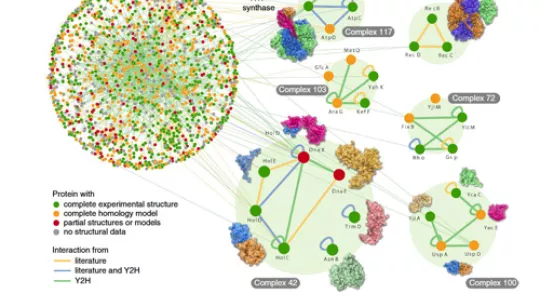
Researchers at the J. Craig Venter Institute and the Virginia Commonwealth University in the United States, in close collaboration with scientists at the Institute for Research in Biomedicine (IRB Barcelona), have published the first map of the molecular interaction of proteins—called the interactome—of Escherichia coli (E. coli).
This map allows researchers to start to understand the intricacies of the bacterial machinery. The study reveals around 25% (2,234) of the approx. 10,000 key interactions estimated for E. coli and covers about 70% of its proteome (the collection of proteins).
The study is published in Nature Biotechnology, one of the journals of reference for the scientific community, and its of high relevance due to the fundamental role that E. coli plays in white biotechnology, that is to say, for the production of large amounts of chemical material, such as artemisinin to treat malaria and insulin for diabetes.
The genome of E. coli is widely known—tens of strains have been sequenced, ranging from some that cause disease to others that do not—as is the proteomics of complexes (map of protein complexes), and it is the most used model organism for metabolomic (the map of metabolic cycles) research.
The information obtained from these large-scale –omics studies has allowed researchers to generate a list of the molecular components of E. coli that are involved in its function. However, in order to have a complete map of an organism, of how it works, it was necessary to piece together the interactome, the molecular interactions between the elements described, and particularly between proteins.
“That is what we have done,” says bioinformatician Patrick Aloy. “The map that we present allows us to start to see the intricacies of the bacteria and allows us to design, for example, antibiotics to break the interactions between proteins and to dismantle parts of its molecular machinery or simply understand its entire complexity,” explains the ICREA researcher at IRB.
The “Structural Bioinformatics and Network Biology” group has performed the computational analysis of the interactions networks and has modelled the three-dimensional structures.
“Without the structures you are blind and you cannot design rational strategies to tinker with a complex living system. This study helps us to understand how the E. coli cell networks work and how they control the vital functions of the bacteria”, says Aloy.
The bioinformatics work was undertaken by Roberto Mosca and Arnaud Ceol, postdoctoral fellows in Aloy’s lab, using Interactome3D and NetAligner, methods developed by the group. “Roberto’s and Arnaud’s maps are a key element in the article, since they have allowed the structural annotation and bioinformatic analyses of the interaction networks respectively”, explains Aloy.
The last part of the work has served to confirm “what was already suspected”, explains Aloy, such as why the E. coli bacteria is an ideal model organism, and the conclusions can be extrapolated to many other bacteria.
The study includes a comparison with the proteomes of another 20 bacteria often associated to severe infections, such as those produced by Helicobacter pylori, Vibrio cholera, Haemophilus influenzae, Neisseria meningitides and Yersinia pestis.
“We provide useful information for biotechnology applications, for the understanding of pathologies, and for drug design,” concludes Aloy, who has several collaboration projects with the J. Craig Venter Institute.
The design of new drugs to fight infections is the main objective of AntiPathoGN, an EU project that ended last June and in which Aloy’s group was involved. The results of this study in Nature Biotechnology are also part of this European project, which in coming months will give rise to further results such as therapeutic strategies to disrupt the E. coli interaction network.
Reference article:
The binary protein-protein interaction landscape of Escherichia coli
Seesandra V Rajagopala, Patricia Sikorski, Ashwani Kumar, Roberto Mosca, James Vlasblom, Roland Arnold, Jonathan Franca-Koh, Suman B Pakala, Sadhna Phanse, Arnaud Ceol, Roman Häuser, Gabriella Siszler, Stefan Wuchty, Andrew Emili, Mohan Babu, Patrick Aloy, Rembert Pieper & Peter Uetz.
Nature Biotechnology (2014) doi: 10.1038/nbt.2831
About IRB Barcelona
The Institute for Research in Biomedicine (IRB Barcelona) pursues a society free of disease. To this end, it conducts multidisciplinary research of excellence to cure cancer and other diseases linked to ageing. It establishes technology transfer agreements with the pharmaceutical industry and major hospitals to bring research results closer to society, and organises a range of science outreach activities to engage the public in an open dialogue. IRB Barcelona is an international centre that hosts 400 researchers and more than 30 nationalities. Recognised as a Severo Ochoa Centre of Excellence since 2011, IRB Barcelona is a CERCA centre and member of the Barcelona Institute of Science and Technology (BIST).