Images
Scientists at IRB Barcelona identify a key component of the machinery that allows Staphylococcus aureus to transfer genes that confer antibiotic resistance.
Infection by antibiotic-resistant S. aureus is a serious threat in hospitals worldwide.
Halting the spread of antibiotic-resistant bacteria is one of the strategies available to tackle hospital infections.
Antibiotic resistance of the bacterium Staphylococcus aureus is responsible for 11,300 deaths a year in the United States alone—a figure that corresponds to half of all deaths caused by Gram-positive resistant bacteria in that country. Such high mortality is related to the speed at which the bacterium acquires resistance to antibiotics. A study performed at the Institute for Research in Biomedicine (IRB Barcelona) and involving the collaboration of the Centro de Investigaciones Biológicas (CIB-CSIC) in Madrid has identified a key component of the machinery that S. aureus uses to acquire and transfer genes that confer resistance to antibiotics.The work has been published this week in the Proceedings of the National Academic of Sciences of the United Stated of America (PNAS).
“The battle against bacteria—particularly in the hospital setting where they are a major threat—implies understanding how genes are transferred to adapt to a changing environment. For example, when they are treated with new antibiotics,” explains the head of the study and IRB Barcelona group leader Miquel Coll, also a CSIC researcher, who studies horizontal gene transfer from a structural biology perspective.
Halting the spread
“Horizontal gene transfer confers bacteria with an extraordinary capacity to evolve and adapt rapidly—a capacity that humans do not have for example,” says Coll. One of these pathways is called conjugation, a process by which two bacteria join and one of them transfers a piece of DNA, called plasmid, to the other. “A plasmid is a small piece of circular DNA that holds very few genes, often including those for antibiotic resistance and it takes only a few minutes to be passed between bacteria,” he explains.
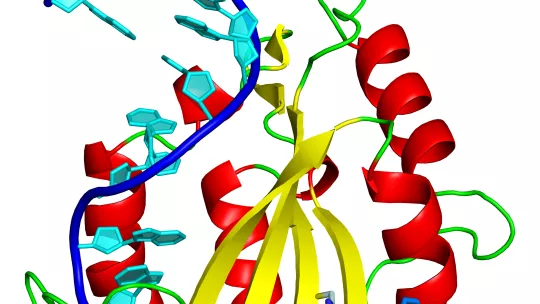
Horizontal gene transfer through conjugation requires molecular machinery, in which an enzymatic protein called relaxase, is a key component. Thanks to the high resolution 3D structure of the complex formed by the relaxase with a fragment of the plasmid DNA, the researchers have identified that an amino acid histidine is a pivotal element in the DNA processing and thus in the transfer and the spread of resistance.
“What we have discovered is that the type of relaxases that are predominant in S. aureus use a DNA processing mechanism that has not been observed so far in other relaxases or any type of known DNA processing proteins,” explains the first author of the study, Radoslaw Pluta, a former “la Caixa” PhD student at IRB Barcelona, and currently a postdoctoral researcher at the International Institute of Molecular and Cell Biology in Warsaw, Poland.
Histidine is the catalytic residue that allows the relaxase to cut DNA, bind to it, stretch one of the two strands and take it into the recipient bacterium, where the relaxase re-join the cut DNA strand, followed by the strand replication to recreate the double strand form of the plasmid DNA again. The recipient bacteria holds now the functional plasmid encoding the resistance genes and the machinery to transfer them to another bacterium. The scientists indicate that this catalytic histidine type of relaxases are present in 85% of Staphylococcus aureus isolates for which genetic information are deposited in the National Center for Biotechnology Information (NCBI; USA).
To test whether histidine is decisive in horizontal gene transfer,, researchers in Manuel Espinosa’s group at the CIB-CSIC, who participated in the study, replaced it by a different amino acid and confirmed that it prevents transfer in culture dishes.
The mutation of histidine does not kill that bacterium but rather prevents gene transfer. How could this mechanism be exploited to fight infections? “I don’t know,” says Coll, “but we now know more details about a lethal bacterium and this may pave the way to the development of molecules to prevent the spread of resistant strains”.
Coll explains that hospital infections are the most difficult types to tackle. “We are in a race that we always lose because when a new antibiotic is brought out, resistance quickly emerges and spreads,” he describes. The scientist adds that the list of antibiotics for hospital use is “too” short. Apart from the difficulty involved in developing new antibiotics, Coll also comments on another obstacle impeding advancement. “There is little investment because the pharmaceutical industry has other priorities. While this is perfectly valid, resources from the public and private sectors should be pooled”.
This work has involved the collaboration of Modesto Orozco’s group, also at IRB Barcelona, which has performed the theoretical studies to validate the chemical reaction between the plasmid DNA and the protein via histidine. The high resolution 3D structure of the protein-DNA complex has been achieved through X-ray crystallography using diffraction data collected at the European Synchrotron Radiation Facility (ESRF) in Grenoble.
Reference article:
Radoslaw Pluta, D. Roeland Boer, Fabián Lorenzo-Díaz, Silvia Russi, Hansel Gómez, Cris Fernández-López, Rosa Pérez-Luque, Modesto Orozco, Manuel Espinosa and Miquel Coll
PNAS (2017): doi: 10.1073/pnas.1702971114
About IRB Barcelona
Created in 2005 by the Generalitat de Catalunya (Government of Catalonia) and University of Barcelona, IRB Barcelona is a Severo Ochoa Centre of Excellence, a seal that was awarded in 2011. The institute is devoted to conducting research of excellence in biomedicine and to transferring results to clinical practice, thus improving people’s quality of life, while simultaneously promoting the training of outstanding researchers, technology transfer, and public communication of science. Its 25 laboratories and seven core facilities address basic questions in biology and are orientated to diseases such as cancer, metastasis, Alzheimer’s, diabetes, and rare conditions. IRB Barcelona is an international centre that hosts 400 employees and 32 nationalities. It is located in the Barcelona Science Park. IRB Barcelona forms part of the Barcelona Institute of Science and Technology (BIST) and the “Xarxa de Centres de Recerca de Catalunya” (CERCA).
About IRB Barcelona
The Institute for Research in Biomedicine (IRB Barcelona) pursues a society free of disease. To this end, it conducts multidisciplinary research of excellence to cure cancer and other diseases linked to ageing. It establishes technology transfer agreements with the pharmaceutical industry and major hospitals to bring research results closer to society, and organises a range of science outreach activities to engage the public in an open dialogue. IRB Barcelona is an international centre that hosts 400 researchers and more than 30 nationalities. Recognised as a Severo Ochoa Centre of Excellence since 2011, IRB Barcelona is a CERCA centre and member of the Barcelona Institute of Science and Technology (BIST).