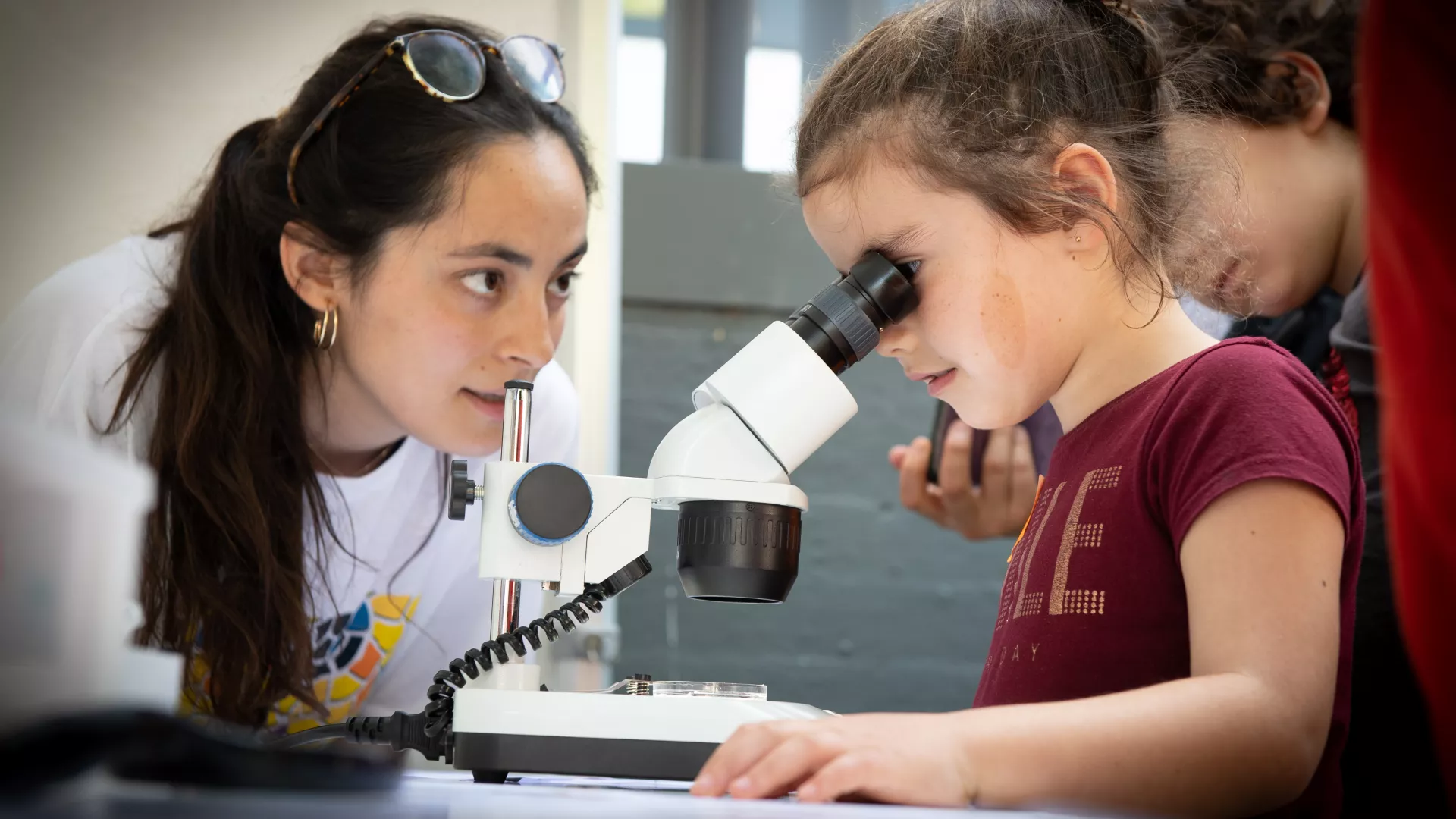
Organized by IRB Barcelona in collaboration with the Fundació Catalunya – La Pedrera.
This year's tutors: Jesús Herráiz, Sylwia Gawrzak, Salvador Pérez, Bahareh Eftekharzdeh, Berta Terré, Paula Martínez-Cristóbal, Carlos Maíllo Carbajo, Julia García, Antonija Kuzmanic, Arturo Rodríguez-Banqueri, Joana Raquel Faria, Mohammed Rahman.
Presentation

Participants of the “Crazy About Biomedicine” 2015 course.
Objectives
Crazy About Biomedicine is a year-long course directed at students in their first year of baccalaureate who wish to explore some of the exciting discoveries being made in the life sciences. Through this course, students will have a chance to deepen their knowledge of scientific theory and techniques in the field of biomedicine. They will work alongside our young researchers to get a taste for what doing science in a top international research institute is like, gain some hands-on experience in the latest cutting-edge methodologies, and position themselves for a potential career in the life sciences.
Course Description
This course includes a combination of theoretical lectures and practical hands-on experimental activities, which take place on 18 Saturdays throughout the year. Presented by IRB Barcelona PhD students and postdocs, the course will cover 12 hot scientific topics, ranging from cell and molecular biology to structural and computational biology, and chemistry. In the first ‘semester’ (Jan-June 2016), the first three Saturdays will be devoted to these general lectures for all participants. During the following six sessions, small groups will enter the labs for hands-on practical experience. This schedule will then be repeated with six new research topics in the second semester (June-Dec). Participants must commit themselves to attending the whole course.
Course language
All lectures and practical sessions will be conducted in English.
Course dates and times
The course will run from January to December 2015, 10.00-14.00h.
SEMESTER I
- Fri. 9 January: Inauguration at Mon Sant Benet- Sant Fruitós del Bages
- Sat. 17 January: Talk 1
- Sat. 31 January: Talk 2
- Sat. 7 February: Talk 3
- Sat. 21 February: Practical session 1
- Sat. 7 March: Practical session 2
- Sat. 21 March: Practical session 3
- Sat. 11 April: Practical session 4
- Sat. 25 April: Practical session 5
- Sat. 9 May: Practical session 6
SEMESTER II
- Sat. 23 May: Talk 1
- Sat. 30 May: Talk 2
- Sat. 13 June: Talk 3
- Sat. 17 September: Practical session 1
- Sat. 3 October: Practical session 2
- Sat. 17 October: Practical session 3
- Sat. 31 October: Practical session 4
- Sat. 14 November: Practical session 5
- Sat. 28 November: Practical session 6
- January 2016: Closing ceremony
Course fees
Foundation Catalunya – La Pedrera will charge 120 euros in terms of administrative and insurance costs (payable directly to the Foundation). The access to the IRB Barcelona facilities and course materials are free of charge. Lunches and snacks are not provided. Students will receive a Certificate of Participation upon successful completion of the course at a special ceremony in January 2016, to which parents and teachers will be invited.
Course location
Institute for Research in Biomedicine (IRB Barcelona)
c/ Baldiri Reixac, 10
Barcelona
Who can apply
This course is directed toward students in the first year of their baccalaureate, who have a special interest and talent in the fields related to the life sciences (primarily biology and chemistry).
Students may apply to a maximum of 2 programmes within the "Crazy About Science" series, and can participate in only one.
How to apply
Interested students must fill in the online application form and include a letter of motivation. A letter of recommendation will be requested directly from two of their teachers, who should know the applicant well. If the applicant has recently changed school then the letters of recommendation should be requested from his/her old teachers. The deadline for registration is 14 October, 2014.
The course is open to a total of 24 students. Candidates will be selected on the basis of their academic record, teacher recommendations and motivation to participate. A short list of candidates will be invited for interviews with the scientific organizers in November after which the final selection will be made. Students will be informed of the outcome by the first week in December. The students selected to participate and their parents/guardians will be asked to sign a letter of commitment to attend all sessions.
Important dates
- 14 October 2014: Application deadline
- Week of 9 November 2014: Short-listed candidates contacted
- 16-27 November 2014: Interviews
- 1-4 December 2014: List of selected students published
- Fri, 9 January 2015: Inauguration at Mon Sant Benet –Sant Fruitós del Bages
- Sat, 17 January 2015: Course begins
Collaborators
Fundació Catalunya La Pedrera, Parc Científic de Barcelona

![]()

If you have any question, please contact us at: irb_outreach@irbbarcelona.org
Programme
SEMESTER 1
1. Having a closer look at peptides
Jesús Herráiz (Albericio’s Lab)
Proteins are large, complex molecules that play many critical roles in our bodies. They do most of the work in cells and are required for the function, defense, and signaling of the body’s tissues and organs. Understanding their structure will allow us to decipher their function, evolutionary development, and interactions.
Proteins are biological molecules that consist of one or more long chains of amino acid residues. A linear chain of such residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than about 20-30 residues, are commonly called peptides. Given their small size, working with these molecules proves difficult.
In this course, we will look at some of the current methods that researchers use to work efficiently with peptides.
The knowledge gained in this course will be use in the course directly below
2. Overview of metastasis: how do cancer cells colonize distant organs?
Sylwia Gawrzak (Gomis’ Lab)
Metastasis is a complex multistep process in which cancer cells leave the original tumor site and migrate to other parts of the body, mainly through the bloodstream. Metastatic relapse, a late event in cancer progression, is associated with poor prognosis. Although strategies targeting primary tumors have been improved, systemic treatments to prevent metastasis are less effective. Consequently, metastasis remains a challenging clinical problem, and the dissection of its biological mechanisms is crucial in order to reveal therapeutic targets to prevent and cure malignant tumors.
This course will give general overview of cancer biology and will focus on the different aspects of metastasis. We will learn about where cancer cells usually form metastases, the essential steps in this process, and the time it takes. The theoretical part will be illustrated by examples of experimental techniques that are used to study metastasis. During the practical part, we will perform histological staining and compare several mouse tissue samples in order to identify metastatic lesions in various organs.
3. A code driving development
Salvador Pérez (Azorín’s Lab)
Eukaryote DNA is not naked but organized in a structure called chromatin, which is formed by nucleosomes. A nucleosome is a particle that comprises core histones, which DNA wraps around. During recent years, our knowledge of chromatin regulation and function has improved through the identification of several histone modifications, histone variants, remodeling complexes, and other structural and non-structural proteins. All these factors participate in chromatin compaction, a process that leads to changes in gene expression. The field of epigenetics studies how these factors affect gene expression and its importance during development, and also how they influence cell processes such as replication and transcription.
This course focuses on the study of the molecular basis of chromatin structure, function and regulation during the cell cycle and the development of an organism. We will use Drosophila melanogaster (fruit flies!) as a model organism. During the course, students will come to realize that human development shares many and considerable similarities with flies.
During this course we will address one of the proteins that binds to DNA and participates in chromatin organization. In this regard, we will concentrate on one of the variants of histone H1 and several modifications, in order to examine how this protein affects gene expression and the development of the fruit fly.
4. An introduction to neurodegeneration and neurodegenerative diseases
Bahareh Eftekharzdeh (Salvatella’s Lab)
Alzheimer’s disease is the most common form of dementia. It has no cure, and symptoms worsen as the disease progresses, eventually leading to death. It was first described by German psychiatrist and neuropathologist Alois Alzheimer in 1906. The causes and progression of Alzheimer's disease are not well understood. Research indicates that this condition is associated with plaques and tangles of amyloid-beta peptide in the brain. Current treatments alleviate the symptoms of the disease, but none stop or reverse its progression. As of 2012, more than 1000 clinical trials have been conducted to find ways to treat Alzheimer’s, but whether any of the treatments being tested will be effective remains to be seen. While mental stimulation, exercise, and a balanced diet have been proposed as ways to delay cognitive symptoms (though not brain pathology) in healthy elderly individuals, there is no conclusive evidence supporting an effect. This course will take an in-depth look at the mechanisms of neurodegeneration, with a focus on Alzheimer’s disease and Parkinson’s disease.
We will begin with an introduction to the history of these diseases, looking at when they were first detected in the brains of patients. This will be followed by an explanation of the molecular and cellular mechanisms involved. We will then go through the hypotheses of the causes, and the latest research strategies to find cures. Finally, we will finish the course with a question and answer session in which students will have the opportunity to reflect on future approaches in this field of research.
5. Understanding DNA: What makes you different from everybody else?
Berta Terré (Stracker’s Lab)
DNA is the genetic “instruction manual” found in all our cells. This molecule contains genetic information that controls cellular functions such as proliferation and apoptosis, and even eye color. However, sometimes DNA can undergo modifications that can disrupt these biological processes, thus leading to the development of diseases such as cancer. Our cells are equipped with a system to control and repair these lesions, namely the DNA Damage Response pathway. Simply put, some genes serve as guardians of the genome, being responsible for maintaining genomic stability and protecting cells. However, this system does not work in exactly the same way for everyone, and this is why, despite having similar lifestyles, some people develop cancer while others do not.
In this two-part course, we will explore the association between genomic instability and cancer. We will address the mechanisms by which DNA controls cellular function and how the genetic differences between people influence their risk of getting cancer.
This course will be continued in Semester II.
6. Why mitochondrial dynamic is essential for live
Paula Martínez-Cristóbal (Zorzano’s Lab)
Mitochondria are mobile organelles present in dynamic networks. They continuously join by fusion and divide by fission. The secret of the dynamics of mitochondria is revealed by confocal live-cell imaging with the use of potentiometric dyes or mitochondria-targeted fluorescent proteins.
Fission and fusion regulate fundamental cellular processes, such as calcium homeostasis and the generation of ATP and reactive oxygen species, and consequently, they have important roles in cell-cycle progression, apoptosis, mitophagy, and oxygen sensing. However, the disruption of mitochondrial dynamics contributes to the pathogenesis of complex diseases that are not considered to involve mitochondria; these diseases include cancer, cardiovascular disease, and neurodegenerative diseases.
In this course, we will explore the relationship between mitochondrial dynamics, apoptosis, and autophagy. The students will perform experiments with cells and will work on elucidating the role of some mitochondrial proteins actively involved in mitochondrial fusion.
SEMESTER 2
1. Regulation of gene expression or why we don't understand our genome (yet)
Carlos Maíllo Carbajo (Mendez’ lab)
In the year 2000 the Human Genome Project was completed at a cost of around 3 billion dollars and ten years of intense work. The main conclusion was that over 98% of our genetic material was junk DNA. Far from resolving our questions, reading our genome made us recognize we didn't understand it. Since then, mounting evidence has made us realize that to understand our genome means to understand how its expression is regulated.
Now we are starting to get a glimpse of how the cells counts on extremely complex regulatory networks to switch genes on and off through a myriad of novel players such as microRNAs, long non-codingRNAs and RNA-binding proteins.
Understanding how gene expression is controlled will be vital to know how an erythrocyte and a neuron are so different while sharing the exact same genes or why and individual gets a disease like cancer while and identical twin remains completely healthy.
In this course we will take a look at the current methods that researchers use to study the regulation of gene expression using in vitro cell cultured systems.
2. Designing peptide drugs
Júlia Garcia (Giralt’s Lab)
Peptides are small proteins made up of a small number of amino acids. The distinction between proteins and peptides is somewhat arbitrary, although proteins typically contain 50 or more amino acids, whereas peptides contain fewer.
Natural peptides are present in our organism, where they play several biological roles. The main ones involve interaction with macromolecules such as proteins. For this reason, peptides can be used as therapeutic agents to treat a number of diseases. These potential drugs can be chemically synthesized using a method called solid-phase peptide synthesis (SPPS). While this technique offers numerous advantages, it is crucial that it be designed properly for each specific case.
Since most biological targets are usually inside cells, peptides need to be able to cross cell membranes, which act as barriers. Certain types of peptides, such as some cyclic or bicyclic molecules, have the capacity to cross these membranes when specific elements are introduced into their sequences. In the practical session, we will attempt to design and synthesize a therapeutic peptide.
3. Rational drug design
Antonija Kuzmanic (Orozco’s Lab)
The capacity of X-ray crystallography to determine protein structures has provided scientists with an innovative way to find new drugs. By combining this approach with emerging computational methods, rational drug design not only has significantly reduced the costs of this process, but has made the treatment of many diseases possible, even on a personalized level.
This part of the course seeks to teach students how to combine structural information with computational methods, such as molecular docking, in order to discover a novel drug. Students will be given a list of proteins that are successful examples of rational drug design and will be asked to choose one that interests them. We will then study the basics of the protein structure, the features of its binding pockets, and how the possible ligands for these pockets are selected, and we will use docking software to evaluate whether the proposed compounds will make effective drugs.
4. Proteins in biological membranes. What do they look like?
Arturo Rodríguez-Banqueri (Palacin’s Lab)
It has been estimated that between 20 and 30% of the proteomes of most organisms are membrane proteins. Membrane proteins can perform a large variety of relevant functions, being key players in maintaining cell homeostasis by transferring information between the extracellular and intracellular spaces of the cell or between intracellular compartments. It is not surprising, therefore, that the malfunction of these proteins is a direct cause of disease. In fact, almost 50% of current drugs target membrane proteins, thus reflecting the relevance of these molecules in in drug discovery and development.
X-ray crystallography is currently one of the most powerful techniques to determine protein structure at the atomic level. Information on the structure of these molecules is essential for drug design. Unfortunately, obtaining high quality crystals of membrane proteins for X-ray diffraction is difficult because of the hydrophobic nature of the proteins. Their low stability in solution and their tendency to form aggregates are the biggest problems encountered during crystallization studies.
In this course, we will take a look at the protein engineering techniques that researchers use to work with membrane proteins. This stage is crucial to accomplish successful crystallization trials and to obtain atomic-level resolution of a protein structure performing X-ray crystallography.
5. Understanding DNA: What makes you different from everybody else?
This course is a continuation from Semester I.
Joana Raquel Faria (Stracker’s Lab)
6. Stem cells, disease modeling and drug discovery using… flies?
Mahi Rahman (Casanova’s Lab)
Drosophila is a beautiful flying insect that has been used to study the basic biology of life at the cellular and genetic level for over a century. With a short lifespan, only four chromosomes, and devoid of multiple isoforms of genes for the most part, this invertebrate has become a very useful and efficient tool to interrogate complex biological issues, ranging from genetic mutations to the environmental effects on the health of organisms. The discovery of intestinal stem cells in 2006 also unleashed a very interesting frontier to explore stem cell-derived cancer—a hallmark of relapse for many types of cancer. In addition, as we can model many aspects of cancer biology, this invertebrate is a smart choice for screening drugs, a process that is prohibitively expensive in higher vertebrates.
In this course, we will start with a brief history of Drosophila as the workhorse for biological studies and the advantages and limitations of this paradigm. We will study the intestinal stem cells in Drosophila and how diseases such as colorectal cancer are modeled. In addition, we will explore how Drosophila is used for drug discovery purposes. In the process, we will also learn how to design a scientific project, work with Drosophila, dissect and identify various cell types, and use advanced imaging techniques such as confocal microscopy. Finally, we will learn about the latest frontiers of biomedical research with respect to this model.
Venue
CRAZY ABOUT BIOMEDICINE will take place at the IRB Barcelona facilities.
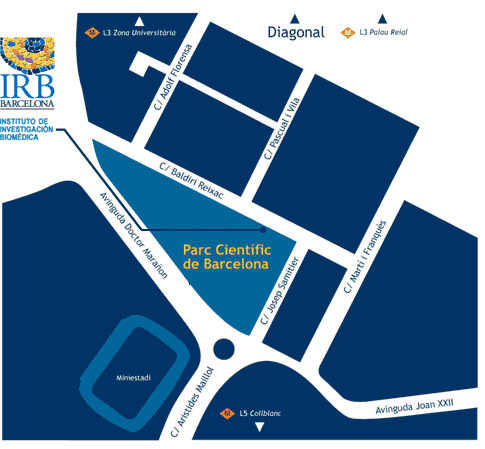
Venue
IRB Barcelona
c/o Parc Cientific de Barcelona
Carrer Baldiri Reixac, 10
08028 Barcelona
(Campus de la Diagonal, Universitat de Barcelona)

