Protein Expression
The Protein Expression Core Facility (PECF) accelerates IRB research by providing advanced technologies for cloning, expression, and purification of vital proteins, with capabilities for high-throughput (HTP) processing to enhance efficiency. Established in 2007, PECF offers expertise across the entire gene-to-protein pipeline, utilizing innovative plasmid suites like pOPIN and pPEU for rapid and efficient protein expression and purification.
PECF supports diverse protein expression needs, including prokaryotic systems for producing stable, soluble proteins and hetero-dimers, as well as specialized labeling for structural studies. For more complex proteins, the facility leverages mammalian hosts like 293-6E cells to produce bioactive, glycosylated proteins in high yields. PECF also utilizes insect cells and modular systems like GoldenBac for generating multi-protein complexes. Alongside these services, PECF provides high-quality reagents and offers training for researchers seeking hands-on experience.
For further details or enquiries please contact us (proexp@irbbarcelona.org).

The PECF offers a wide variety of services. For more information please contact proexp@irbbarcelona.org.
- Molecular biology services
- Gene Cloning into pOPIN, pPEU vectors
- Cloning into custom (user-derived) vectors
- Production of (multiples) point mutations
- Production of GoldenBac co-expression transfer plasmids
- Expression Screening services
- P1 baculovirus generation using pOPIN/pPEU/pGB vectors
- P1 baculovirus generation from pFastBac vectors
- Baculovirus amplification in Sf9 cells (P2 to P3)
- Expression screening in Sf9 cells and serum-free HEK cells (His/Strep-tagged proteins)
- Expression and Purification services
- E.coli Expression screening GST/His/Strep-Tagged Proteins in E.coli
- E.coli Expression and Purification of His-, Strep-, or GST- Tagged Proteins at Scale
- Serum-Free Mammalian (HEK293-6E) Expression without protein purification
- Serum-Free Mammalian (HEK293-6E) Expression and Purification of Intracellular His-Tagged Proteins at Scale
- Serum-Free Mammalian (HEK293-6E) Expression and Purification of secreted and intracellular Strep-Tagged Proteins at Scale
- Additional Serum-Free Mammalian (HEK293-6E) Expression culture (includes plasmid maxiprep) for Mammalian (HEK293-6E) Expression and Purification services
- Cleavage of Tag from Fusion Proteins ≤10 mg of protein
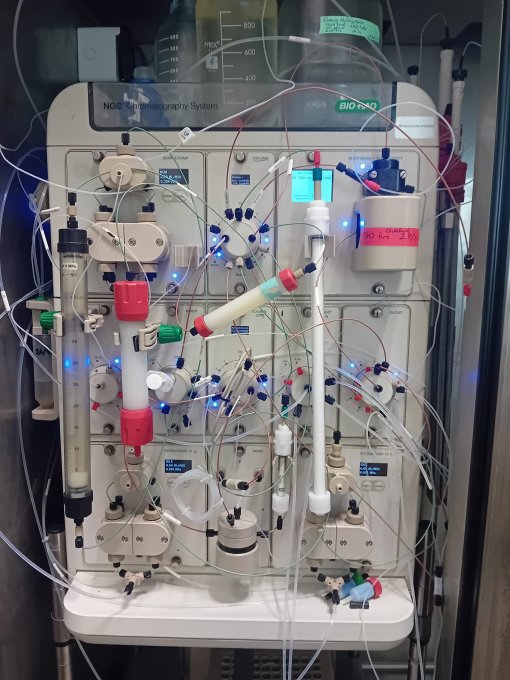
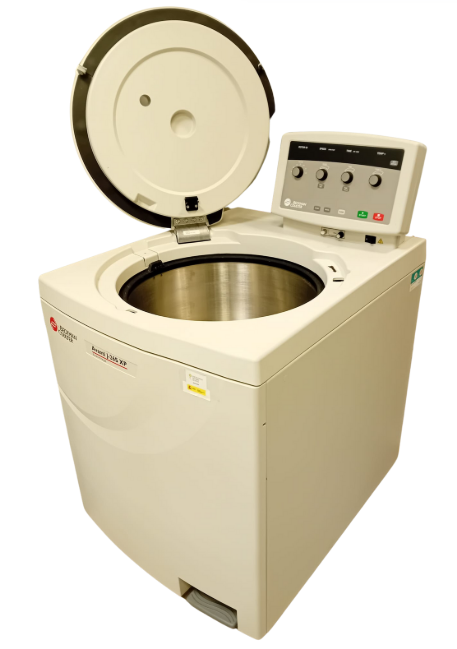



The PECF instrumentation* includes:
- A cell disruptor capable of efficiently lysing cells at 40,000 psi using volumes from a few mls to tens of litres-please contact us if you would like to use this instrument.
- Two NGC FPLC instruments-one of which is capable of fully automated 2D chromatography.
- A fully equipped tissue culture room including a humidified CO2 incubator equipped with three shaker platforms for large-scale mammalian cultures and an epifluorescence inverted microscope for the visualization of expressed fluorescent (fusion) proteins.
- A Nanodrop ND-1000 and a Nanodrop One microlitre volume UV-Visible light Spectrophotometers.
*-please note that only the cell disruptor and the ND-1000 are currently available for public use.
- Initial inquiry: you can send a mail to proexp@irbbarcelona.org to arrange a meeting to discuss the best approach to meet your research goals.
- Request submission: To submit a project proposal to the PECF, please download and complete the proposal form and send it to proexp@irbbarcelona.org.
- Review & approval: CF staff reviews the request and, if needed, contacts the user for clarification.
- Service processing: full process time ranges between 5 and 15 days depending on the service. If there is an anticipated delay, the PECF will advise the researcher accordingly with a new estimated delivery time.
- Results Delivery
- Follow-up meeting (optional)
An example workflow from our mammalian protein expression platform from gene cloning to purified protein.
We generally have many varied projects in progress at any one time but we try to accommodate any reasonable requests as quickly as we are able (see specific Service descriptions for service timings and deliverables). Please consult us if you have any queries about possible projects or collaborations with the Protein Expression Core Facility.